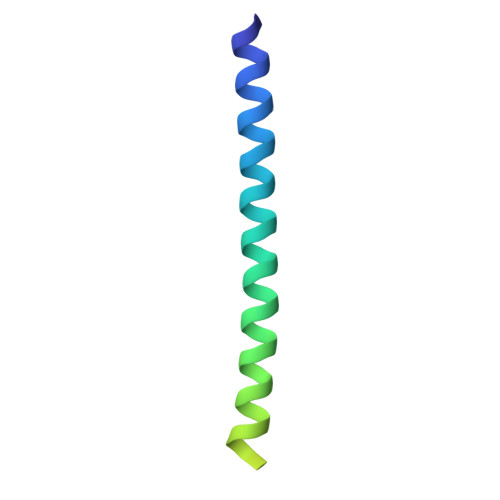
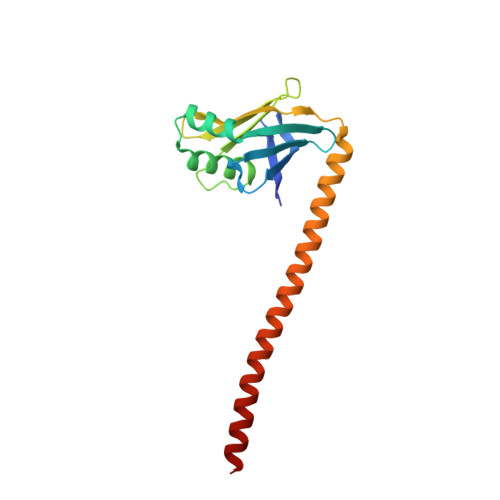
Structure of the tropomyosin overlap complex from chicken smooth muscle: insight into the diversity of N-terminal recognition .
Frye, J., Klenchin, V.A., Rayment, I.(2010) Biochemistry 49: 4908-4920
- PubMed: 20465283 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/bi100349a
- Primary Citation Related Structures:
3MTU, 3MUD - PubMed Abstract:
Tropomyosin is a stereotypical alpha-helical coiled coil that polymerizes to form a filamentous macromolecular assembly that lies on the surface of F-actin. The interaction between the C-terminal and N-terminal segments on adjacent molecules is known as the overlap region. We report here two X-ray structures of the chicken smooth muscle tropomyosin overlap complex. A novel approach was used to stabilize the C-terminal and N-terminal fragments. Globular domains from both the human DNA ligase binding protein XRCC4 and bacteriophage varphi29 scaffolding protein Gp7 were fused to 37 and 28 C-terminal amino acid residues of tropomyosin, respectively, whereas the 29 N-terminal amino acids of tropomyosin were fused to the C-terminal helix bundle of microtubule binding protein EB1. The structures of both the XRCC4 and Gp7 fusion proteins complexed with the N-terminal EB1 fusion contain a very similar helix bundle in the overlap region that encompasses approximately 15 residues. The C-terminal coiled coil opens to allow formation of the helix bundle, which is stabilized by hydrophobic interactions. These structures are similar to that observed in the NMR structure of the rat skeletal overlap complex [Greenfield, N. J., et al. (2006) J. Mol. Biol. 364, 80-96]. The interactions between the N- and C-terminal coiled coils of smooth muscle tropomyosin show significant curvature, which differs somewhat between the two structures and implies flexibility in the overlap complex, at least in solution. This is likely an important attribute that allows tropomyosin to assemble around the actin filaments. These structures provide a molecular explanation for the role of N-acetylation in the assembly of native tropomyosin.
- Department of Biochemistry, University of Wisconsin, Madison, Wisconsin 53706, USA.
Organizational Affiliation: