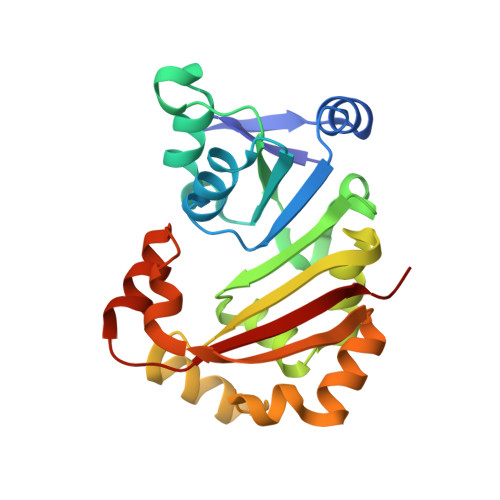
Structural insights into the function of aminoglycoside-resistance A1408 16S rRNA methyltransferases from antibiotic-producing and human pathogenic bacteria.
Macmaster, R., Zelinskaya, N., Savic, M., Rankin, C.R., Conn, G.L.(2010) Nucleic Acids Res 38: 7791-7799
- PubMed: 20639535 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkq627
- Primary Citation Related Structures:
3MQ2, 3MTE - PubMed Abstract:
X-ray crystal structures were determined of the broad-spectrum aminoglycoside-resistance A1408 16S rRNA methyltransferases KamB and NpmA, from the aminoglycoside-producer Streptoalloteichus tenebrarius and human pathogenic Escherichia coli, respectively. Consistent with their common function, both are Class I methyltransferases with additional highly conserved structural motifs that embellish the core SAM-binding fold. In overall structure, the A1408 rRNA methyltransferase were found to be most similar to a second family of Class I methyltransferases of distinct substrate specificity (m(7)G46 tRNA). Critical residues for A1408 rRNA methyltransferase activity were experimentally defined using protein mutagenesis and bacterial growth assays with kanamycin. Essential residues for SAM coenzyme binding and an extended protein surface that likely interacts with the 30S ribosomal subunit were thus revealed. The structures also suggest potential mechanisms of A1408 target nucleotide selection and positioning. We propose that a dynamic extended loop structure that is positioned adjacent to both the bound SAM and a functionally critical structural motif may mediate concerted conformational changes in rRNA and protein that underpin the specificity of target selection and activation of methyltransferase activity. These new structures provide important new insights that may provide a starting point for strategies to inhibit these emerging causes of pathogenic bacterial resistance to aminoglycosides.
- Department of Biochemistry, Emory University School of Medicine, Atlanta, GA 30322, USA.
Organizational Affiliation: