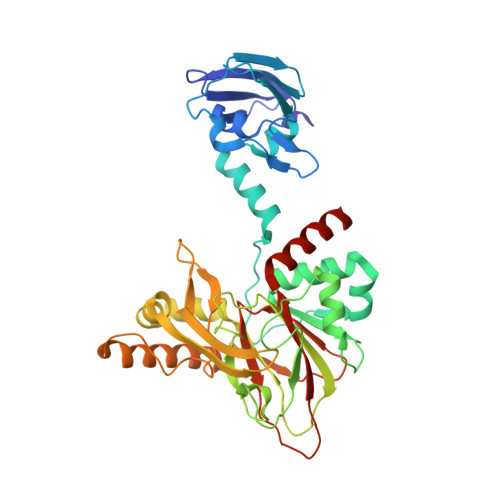
Structural and mechanistic studies on gamma-butyrobetaine hydroxylase.
Leung, I.K., Krojer, T.J., Kochan, G.T., Henry, L., von Delft, F., Claridge, T.D., Oppermann, U., McDonough, M.A., Schofield, C.J.(2010) Chem Biol 17: 1316-1324
- PubMed: 21168767 Search on PubMed
- DOI: https://doi.org/10.1016/j.chembiol.2010.09.016
- Primary Citation Related Structures:
3MS5, 3O2G - PubMed Abstract:
The final step in carnitine biosynthesis is catalyzed by γ-butyrobetaine (γBB) hydroxylase (BBOX), an iron/2-oxoglutarate (2OG) dependent oxygenase. BBOX is inhibited by trimethylhydrazine-propionate (THP), a clinically used compound. We report structural and mechanistic studies on BBOX and its reaction with THP. Crystallographic and sequence analyses reveal that BBOX and trimethyllysine hydroxylase form a subfamily of 2OG oxygenases that dimerize using an N-terminal domain. The crystal structure reveals the active site is enclosed and how THP competes with γBB. THP is a substrate giving formaldehyde (supporting structural links with histone demethylases), dimethylamine, malonic acid semi-aldehyde, and an unexpected product with an additional carbon-carbon bond resulting from N-demethylation coupled to oxidative rearrangement, likely via an unusual radical mechanism. The results provide a basis for development of improved BBOX inhibitors and may inspire the discovery of additional rearrangement reactions.
- The Department of Chemistry and the Oxford Centre for Integrative Systems Biology, Chemistry Research Laboratory, University of Oxford, 12 Mansfield Road, Oxford OX13TA, UK.
Organizational Affiliation: