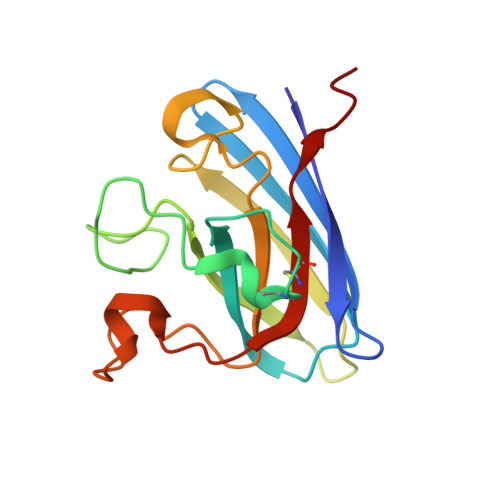
Crystal structure of Cu / Zn superoxide dismutase from Taenia solium reveals metal-mediated self-assembly
Hernandez-Santoyo, A., Landa, A., Gonzalez-Mondragon, E., Pedraza-Escalona, M., Parra-Unda, R., Rodriguez-Romero, A.(2011) FEBS J 278: 3308-3318
- PubMed: 21767346 Search on PubMed
- DOI: https://doi.org/10.1111/j.1742-4658.2011.08247.x
- Primary Citation Related Structures:
3MND - PubMed Abstract:
Taenia solium is the cestode responsible for porcine and human cysticercosis. The ability of this parasite to establish itself in the host is related to its evasion of the immune response and its antioxidant defence system. The latter includes enzymes such as cytosolic Cu/Zn superoxide dismutase. In this article, we describe the crystal structure of a recombinant T. solium Cu/Zn superoxide dismutase, representing the first structure of a protein from this organism. This enzyme shows a different charge distribution at the entrance of the active channel when compared with human Cu/Zn superoxide dismutase, giving it interesting properties that may allow the design of specific inhibitors against this cestode. The overall topology is similar to other superoxide dismutase structures; however, there are several His and Glu residues on the surface of the protein that coordinate metal ions both intra- and intermolecularly. Interestingly, one of these ions, located on the β2 strand, establishes a metal-mediated intermolecular β-β interaction, including a symmetry-related molecule. The factors responsible for the abnormal protein-protein interactions that lead to oligomerization are still unknown; however, high metal levels have been implicated in these phenomena, but exactly how they are involved remains unclear. The present results suggest that this structure could be useful as a model to explain an alternative mechanism of protein aggregation commonly observed in insoluble fibrillar deposits.
- Instituto de Química, Universidad Nacional Autónoma de México, México, DF, México. hersan@servidor.unam.mx
Organizational Affiliation: