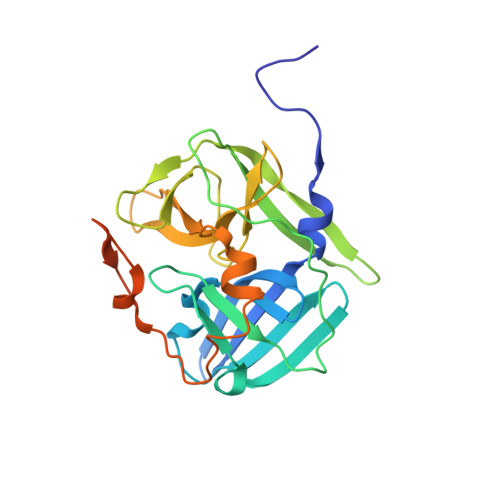
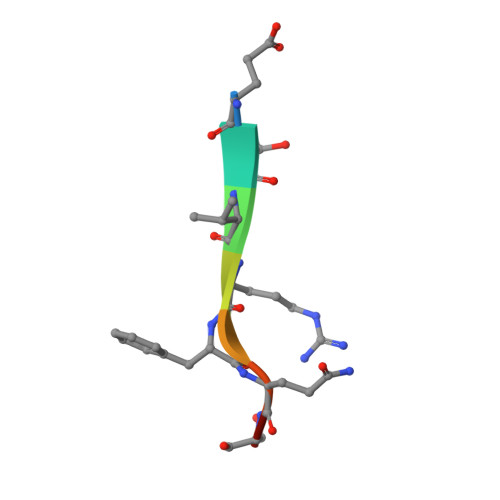
Structural determinants of tobacco vein mottling virus protease substrate specificity.
Sun, P., Austin, B.P., Tozser, J., Waugh, D.S.(2010) Protein Sci 19: 2240-2251
- PubMed: 20862670 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.506
- Primary Citation Related Structures:
3MMG - PubMed Abstract:
Tobacco vein mottling virus (TVMV) is a member of the Potyviridae, one of the largest families of plant viruses. The TVMV genome is translated into a single large polyprotein that is subsequently processed by three virally encoded proteases. Seven of the nine cleavage events are carried out by the NIa protease. Its homolog from the tobacco etch virus (TEV) is a widely used reagent for the removal of affinity tags from recombinant proteins. Although TVMV protease is a close relative of TEV protease, they exhibit distinct sequence specificities. We report here the crystal structure of a catalytically inactive mutant TVMV protease (K65A/K67A/C151A) in complex with a canonical peptide substrate (Ac-RETVRFQSD) at 1.7-Å resolution. As observed in several crystal structures of TEV protease, the C-terminus (∼20 residues) of TVMV protease is disordered. Unexpectedly, although deleting the disordered residues from TEV protease reduces its catalytic activity by ∼10-fold, an analogous truncation mutant of TVMV protease is significantly more active. Comparison of the structures of TEV and TVMV protease in complex with their respective canonical substrate peptides reveals that the S3 and S4 pockets are mainly responsible for the differing substrate specificities. The structure of TVMV protease suggests that it is less tolerant of variation at the P1' position than TEV protease. This conjecture was confirmed experimentally by determining kinetic parameters k(cat) and K(m) for a series of oligopeptide substrates. Also, as predicted by the cocrystal structure, we confirm that substitutions in the P6 position are more readily tolerated by TVMV than TEV protease.
- Macromolecular Crystallography Laboratory, Center for Cancer Research, National Cancer Institute at Frederick, Frederick, Maryland, USA.
Organizational Affiliation: