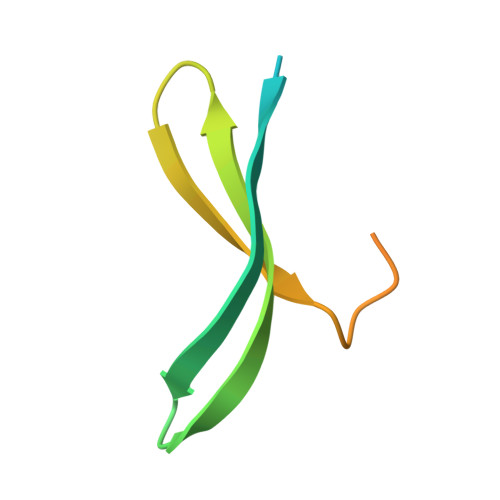
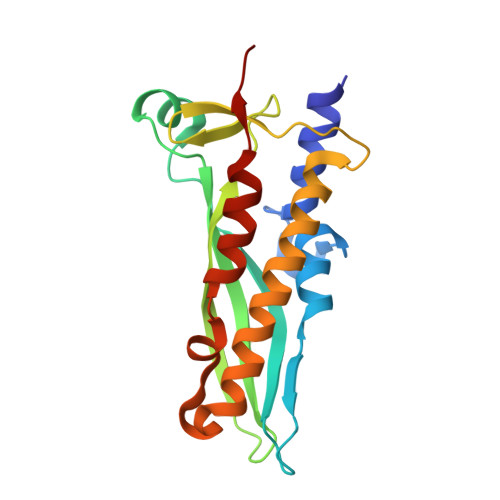
Structural basis for the bacterial transcription-repair coupling factor/RNA polymerase interaction.
Westblade, L.F., Campbell, E.A., Pukhrambam, C., Padovan, J.C., Nickels, B.E., Lamour, V., Darst, S.A.(2010) Nucleic Acids Res 38: 8357-8369
- PubMed: 20702425 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkq692
- Primary Citation Related Structures:
3MLQ - PubMed Abstract:
The transcription-repair coupling factor (TRCF, the product of the mfd gene) is a widely conserved bacterial protein that mediates transcription-coupled DNA repair. TRCF uses its ATP-dependent DNA translocase activity to remove transcription complexes stalled at sites of DNA damage, and stimulates repair by recruiting components of the nucleotide excision repair pathway to the site. A protein/protein interaction between TRCF and the β-subunit of RNA polymerase (RNAP) is essential for TRCF function. CarD (also called CdnL), an essential regulator of rRNA transcription in Mycobacterium tuberculosis, shares a homologous RNAP interacting domain with TRCF and also interacts with the RNAP β-subunit. We determined the 2.9-Å resolution X-ray crystal structure of the RNAP interacting domain of TRCF complexed with the RNAP-β1 domain, which harbors the TRCF interaction determinants. The structure reveals details of the TRCF/RNAP protein/protein interface, providing a basis for the design and interpretation of experiments probing TRCF, and by homology CarD, function and interactions with the RNAP.
- Laboratory of Molecular Biophysics, The Rockefeller University, 1230 York Avenue, New York, NY 10065, USA.
Organizational Affiliation: