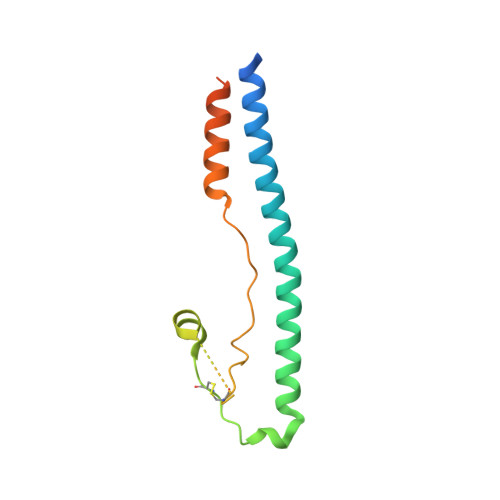
X-ray structure of the arenavirus glycoprotein GP2 in its postfusion hairpin conformation
Igonet, S., Vaney, M.C., Vonhrein, C., Bricogne, G., Stura, E.A., Hengartner, H., Eschli, B., Rey, F.A.(2011) Proc Natl Acad Sci U S A 108: 19967-19972
- PubMed: 22123988 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1108910108
- Primary Citation Related Structures:
3MKO - PubMed Abstract:
Arenaviruses are important agents of zoonotic disease worldwide. The virions expose a tripartite envelope glycoprotein complex at their surface, formed by the glycoprotein subunits GP1, GP2 and the stable signal peptide. This complex is responsible for binding to target cells and for the subsequent fusion of viral and host-cell membranes for entry. During this process, the acidic environment of the endosome triggers a fusogenic conformational change in the transmembrane GP2 subunit of the complex. We report here the crystal structure of the recombinant GP2 ectodomain of the lymphocytic choriomeningitis virus, the arenavirus type species, at 1.8-Å resolution. The structure shows the characteristic trimeric coiled coil present in class I viral fusion proteins, with a central stutter that allows a close structural alignment with most of the available structures of class I and III viral fusion proteins. The structure further shows a number of intrachain salt bridges stabilizing the postfusion hairpin conformation, one of which involves an aspartic acid that appears released from a critical interaction with the stable signal peptide upon low pH activation.
- Département de Virologie, Unité de Virologie Structurale, Institut Pasteur, F-75724 Paris Cedex 15, France.
Organizational Affiliation: