Cyclic amide bioisosterism: Strategic application to the design and synthesis of HCV NS5B polymerase inhibitors.
Yang, H., Hendricks, R.T., Arora, N., Nitzan, D., Yee, C., Lucas, M.C., Yang, Y., Fung, A., Rajyaguru, S., Harris, S.F., Leveque, V.J., Hang, J.Q., Pogam, S.L., Reuter, D., Tavares, G.A.(2010) Bioorg Med Chem Lett 20: 4614-4619
- PubMed: 20584604 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmcl.2010.06.008
- Primary Citation Related Structures:
3MF5 - PubMed Abstract:
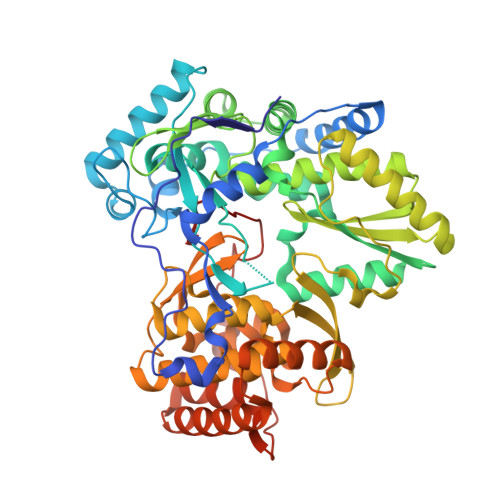
Conformational modeling has been successfully applied to the design of cyclic bioisosteres used to replace a conformationally rigid amide bond in a series of thiophene carboxylate inhibitors of HCV NS5B polymerase. Select compounds were equipotent with the original amide series. Single-point mutant binding studies, in combination with inhibition structure-activity relationships, suggest this new series interacts at the Thumb-II domain of NS5B. Inhibitor binding at the Thumb-II site was ultimately confirmed by solving a crystal structure of 8b complexed with NS5B.
- Medicinal Chemistry Department, Roche Palo Alto, LLC, Palo Alto, CA 94304, USA. Hanbiaoyang@gmail.com
Organizational Affiliation: