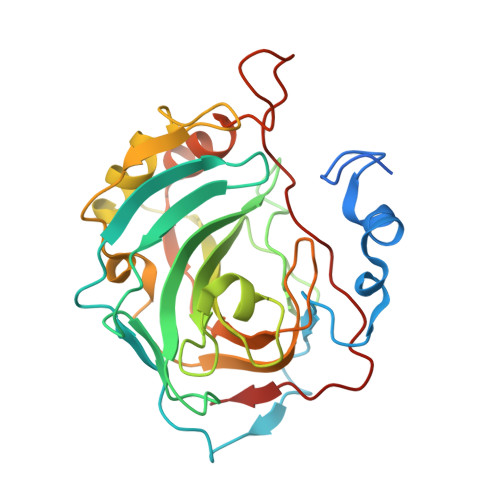
Crystal Structure of Human Carbonic Anhydrase VII [isoform 1], CA7
Ugochukwu, E., Shafqat, N., Pilka, E., Chaikuad, A., Krojer, T., Muniz, J., Kim, J., Bray, J., Bountra, C., Arrowsmith, C.H., Weigelt, J., Edwards, A., von Delft, F., Carpenter, E.P., Yue, W.W., Oppermann, U.To be published.