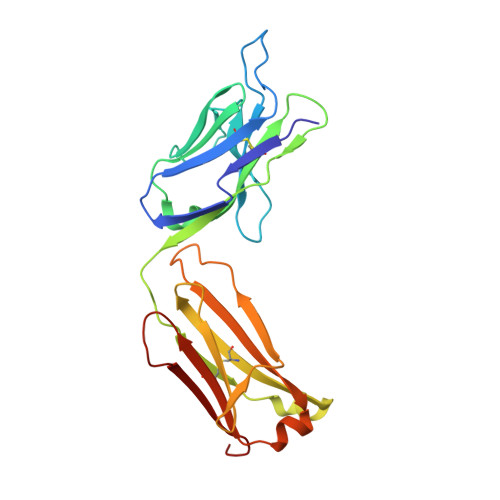
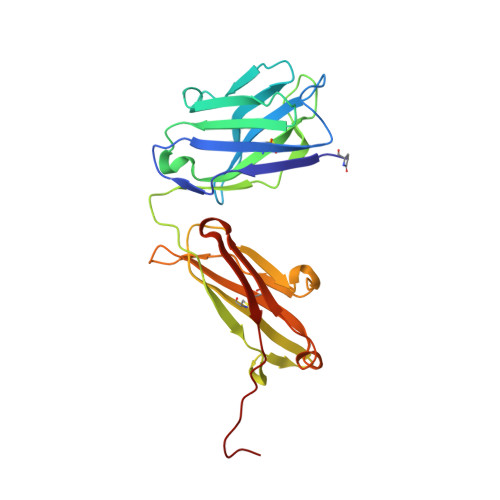
On the domain pairing in chimeric antibodies.
Teplyakov, A., Obmolova, G., Carton, J.M., Gao, W., Zhao, Y., Gilliland, G.L.(2010) Mol Immunol 47: 2422-2426
- PubMed: 20554002 Search on PubMed
- DOI: https://doi.org/10.1016/j.molimm.2010.05.002
- Primary Citation Related Structures:
3MBX - PubMed Abstract:
A chimeric antibody was constructed from two unrelated antibodies by combining their heavy and light chains. The "double chimera" consists of the mouse variable regions of different specificity (IL-13 and EMMPRIN) and the constant regions of different origin (mouse and human). The Fab fragment of this chimeric antibody was expressed in mammalian cells, and the crystal structure was determined at 1.6A resolution. Despite a large number of amino acid substitutions in the double chimera with respect to the parent antibodies, the heavy and light chains associate into a stable molecule. Comparison to the structure of one of the parent antibodies reveals that the variable domain interface, as well as the conformation of antigen-binding loops, is preserved without major rearrangements due to conservation of amino acids in key positions. Comparison to the structures of the all-human and all-mouse constant domains indicates a remarkable plasticity of the inter-chain interface that can tolerate residue relocations of up to 6A.
- Centocor R&D, Inc., 145 King of Prussia Road, Radnor, PA 19087, USA. ateplyak@its.jnj.com
Organizational Affiliation: