Structural Basis of HIV-1 Neutralization by Affinity Matured Fabs Directed against the Internal Trimeric Coiled-Coil of gp41.
Gustchina, E., Li, M., Louis, J.M., Anderson, D.E., Lloyd, J., Frisch, C., Bewley, C.A., Gustchina, A., Wlodawer, A., Clore, G.M.(2010) PLoS Pathog 6: e1001182-e1001182
- PubMed: 21085615 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.ppat.1001182
- Primary Citation Related Structures:
3MA9, 3MAC - PubMed Abstract:
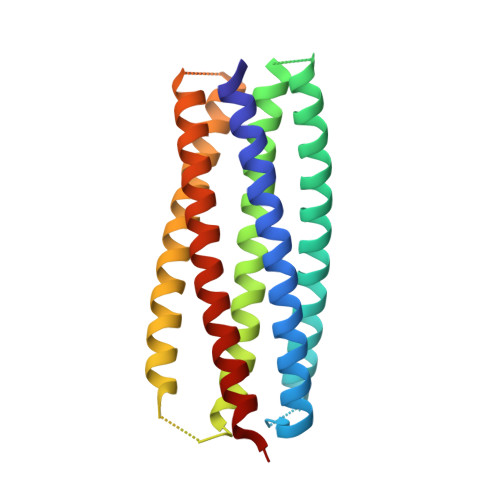
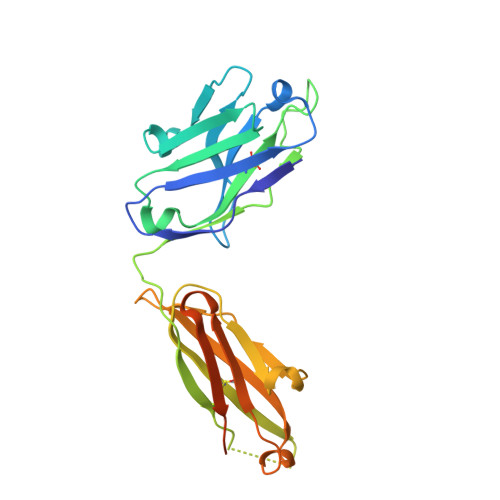
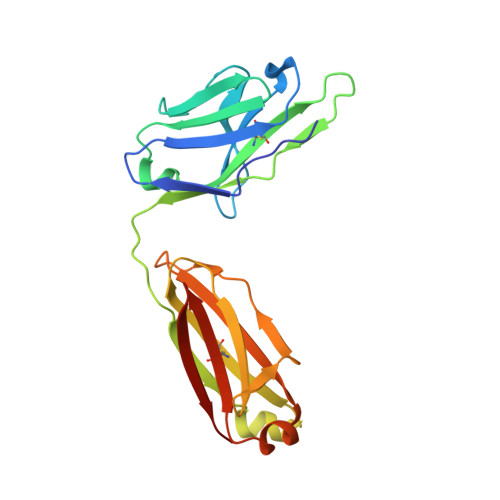
The conserved internal trimeric coiled-coil of the N-heptad repeat (N-HR) of HIV-1 gp41 is transiently exposed during the fusion process by forming a pre-hairpin intermediate, thus representing an attractive target for the design of fusion inhibitors and neutralizing antibodies. In previous studies we reported a series of broadly neutralizing mini-antibodies derived from a synthetic naïve human combinatorial antibody library by panning against a mimetic of the trimeric N-HR coiled coil, followed by affinity maturation using targeted diversification of the CDR-H2 loop. Here we report crystal structures of the N-HR mimetic 5-Helix with two Fabs that represent the extremes of this series: Fab 8066 is broadly neutralizing across a wide panel of B and C type HIV-1 viruses, whereas Fab 8062 is non-neutralizing. The crystal structures reveal important differences in the conformations of the CDR-H2 loops in the complexes that propagate into other regions of the antigen-antibody interface, and suggest that both neutralization properties and affinity for the target can be attributed, at least in part, to the differences in the interactions of the CDR-H2 loops with the antigen. Furthermore, modeling of the complex of an N-HR trimer with three Fabs suggests that the CDR-H2 loop may be involved in close intermolecular contacts between neighboring antibody molecules, and that such contacts may hinder the formation of complexes between the N-HR trimer and more than one antibody molecule depending on the conformation of the bound CDR-H2 loop which is defined by its interactions with antigen. Comparison with the crystal structure of the complex of 5-Helix with another neutralizing monoclonal antibody known as D5, derived using an entirely different antibody library and panning procedure, reveals remarkable convergence in the optimal sequence and conformation of the CDR-H2 loop.
- Laboratory of Chemical Physics, National Institute of Diabetes and Digestive and Kidney Diseases, National Institutes of Health, Bethesda, Maryland, United States of America. gustchia@mail.nih.gov
Organizational Affiliation: