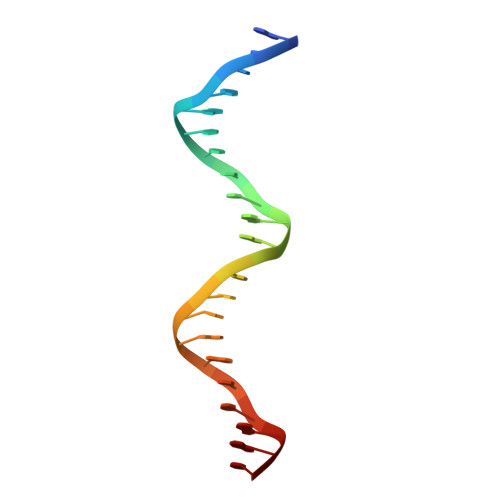
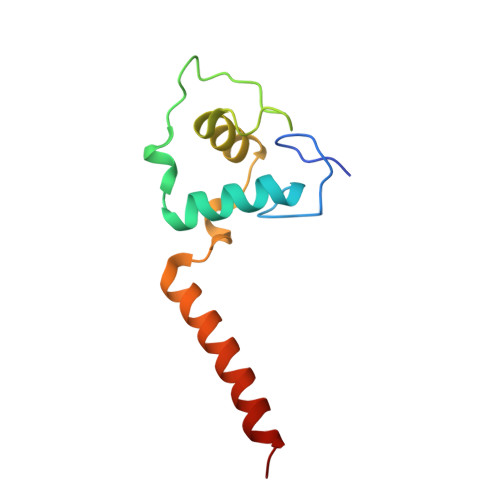
Structure of a thyroid hormone receptor DNA-binding domain homodimer bound to an inverted palindrome DNA response element.
Chen, Y., Young, M.A.(2010) Mol Endocrinol 24: 1650-1664
- PubMed: 20610536 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1210/me.2010-0129
- Primary Citation Related Structures:
3M9E - PubMed Abstract:
Thyroid hormone receptor (TR), as a member of the nuclear hormone receptor family, can recognize and bind different classes of DNA response element targets as either a monomer, a homooligomer, or a heterooligomer. We report here the first crystal structure of a homodimer TR DNA-binding domain (DBD) in complex with an inverted repeat class of thyroid response element (TRE). The structure shows a nearly symmetric structure of the TR DBD assembled on the F2 TRE where the base recognition contacts in the homodimer DNA complex are conserved relative to the previously published structure of a TR-9-cis-retinoic acid receptor heterodimer DNA complex. The new structure also reveals that the T-box region of the DBD can function as a structural hinge that enables a large degree of flexibility in the position of the C-terminal extension helix that connects the DBD to the ligand-binding domain. Although the isolated TR DBDs exist as monomers in solution, we have measured highly cooperative binding of the two TR DBD subunits onto the inverted repeat DNA sequence. This suggests that elements of the DBD can influence the specific TR oligomerization at target genes, and it is not just interactions between the ligand-binding domains that are responsible for TR oligomerization at target genes. Mutational analysis shows that intersubunit contacts at the DBD C terminus account for some, but not all, of the cooperative homodimer TR binding to the inverted repeat class TRE.
- Department of Biophysics, University of Michigan, Ann Arbor, Michigan 48109, USA.
Organizational Affiliation: