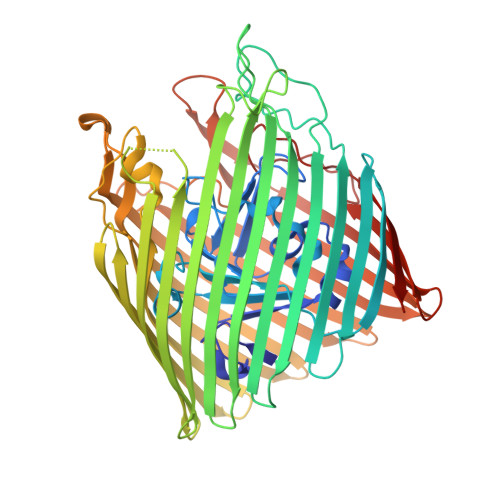
Conformational exchange in a membrane transport protein is altered in protein crystals.
Freed, D.M., Horanyi, P.S., Wiener, M.C., Cafiso, D.S.(2010) Biophys J 99: 1604-1610
- PubMed: 20816073 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.bpj.2010.06.026
- Primary Citation Related Structures:
3M8B, 3M8D - PubMed Abstract:
Successful macromolecular crystallography requires solution conditions that may alter the conformational sampling of a macromolecule. Here, site-directed spin labeling is used to examine a conformational equilibrium within BtuB, the Escherichia coli outer membrane transporter for vitamin B(12). Electron paramagnetic resonance (EPR) spectra from a spin label placed within the N-terminal energy coupling motif (Ton box) of BtuB indicate that this segment is in equilibrium between folded and unfolded forms. In bilayers, substrate binding shifts this equilibrium toward the unfolded form; however, EPR spectra from this same spin-labeled mutant indicate that this unfolding transition is blocked in protein crystals. Moreover, crystal structures of this spin-labeled mutant are consistent with the EPR result. When the free energy difference between substates is estimated from the EPR spectra, the crystal environment is found to alter this energy by 3 kcal/mol when compared to the bilayer state. Approximately half of this energy change is due to solutes or osmolytes in the crystallization buffer, and the remainder is contributed by the crystal lattice. These data provide a quantitative measure of how a conformational equilibrium in BtuB is modified in the crystal environment, and suggest that more-compact, less-hydrated substates will be favored in protein crystals.
- Departments of Chemistry, University of Virginia, Charlottesville, Virginia, USA.
Organizational Affiliation: