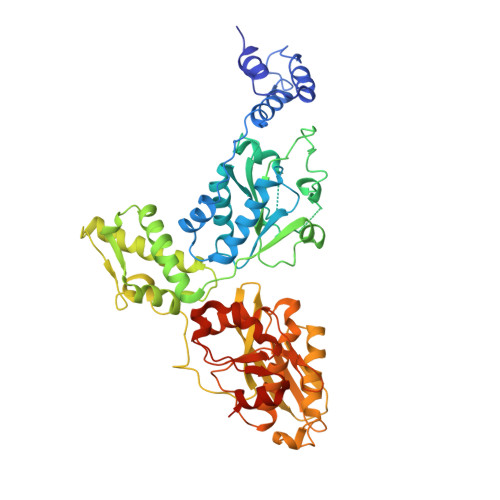
Crystal Structures of Bacillus subtilis Lon Protease.
Duman, R.E., Lowe, J.(2010) J Mol Biology 401: 653-670
- PubMed: 20600124 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2010.06.030
- Primary Citation Related Structures:
3M65, 3M6A - PubMed Abstract:
Lon ATP-dependent proteases are key components of the protein quality control systems of bacterial cells and eukaryotic organelles. Eubacterial Lon proteases contain an N-terminal domain, an ATPase domain, and a protease domain, all in one polypeptide chain. The N-terminal domain is thought to be involved in substrate recognition, the ATPase domain in substrate unfolding and translocation into the protease chamber, and the protease domain in the hydrolysis of polypeptides into small peptide fragments. Like other AAA+ ATPases and self-compartmentalising proteases, Lon functions as an oligomeric complex, although the subunit stoichiometry is currently unclear. Here, we present crystal structures of truncated versions of Lon protease from Bacillus subtilis (BsLon), which reveal previously unknown architectural features of Lon complexes. Our analytical ultracentrifugation and electron microscopy show different oligomerisation of Lon proteases from two different bacterial species, Aquifex aeolicus and B. subtilis. The structure of BsLon-AP shows a hexameric complex consisting of a small part of the N-terminal domain, the ATPase, and protease domains. The structure shows the approximate arrangement of the three functional domains of Lon. It also reveals a resemblance between the architecture of Lon proteases and the bacterial proteasome-like protease HslUV. Our second structure, BsLon-N, represents the first 209 amino acids of the N-terminal domain of BsLon and consists of a globular domain, similar in structure to the E. coli Lon N-terminal domain, and an additional four-helix bundle, which is part of a predicted coiled-coil region. An unexpected dimeric interaction between BsLon-N monomers reveals the possibility that Lon complexes may be stabilised by coiled-coil interactions between neighbouring N-terminal domains. Together, BsLon-N and BsLon-AP are 36 amino acids short of offering a complete picture of a full-length Lon protease.
- MRC Laboratory of Molecular Biology, Hills Road, Cambridge CB2 0QH, UK.
Organizational Affiliation: