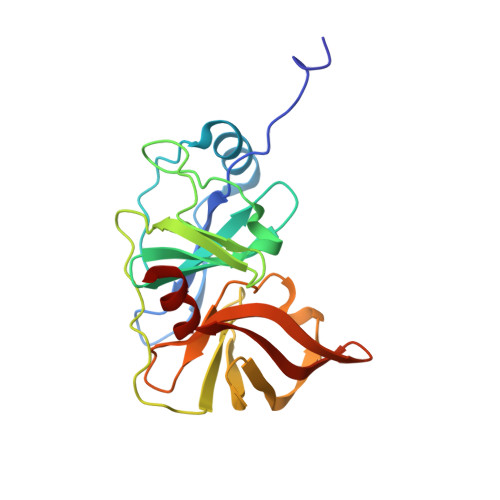
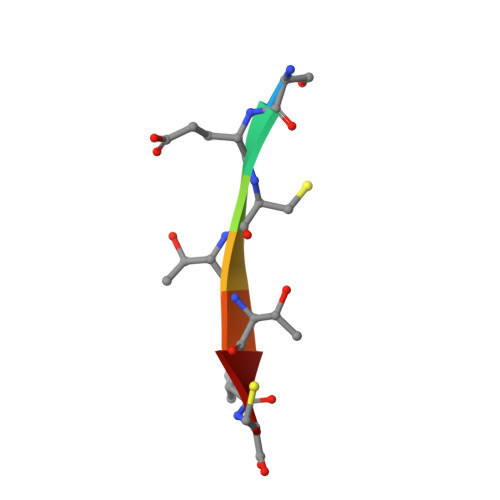
Drug resistance against HCV NS3/4A inhibitors is defined by the balance of substrate recognition versus inhibitor binding.
Romano, K.P., Ali, A., Royer, W.E., Schiffer, C.A.(2010) Proc Natl Acad Sci U S A 107: 20986-20991
- PubMed: 21084633 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1006370107
- Primary Citation Related Structures:
3M5L, 3M5M, 3M5N, 3M5O - PubMed Abstract:
Hepatitis C virus infects an estimated 180 million people worldwide, prompting enormous efforts to develop inhibitors targeting the essential NS3/4A protease. Resistance against the most promising protease inhibitors, telaprevir, boceprevir, and ITMN-191, has emerged in clinical trials. In this study, crystal structures of the NS3/4A protease domain reveal that viral substrates bind to the protease active site in a conserved manner defining a consensus volume, or substrate envelope. Mutations that confer the most severe resistance in the clinic occur where the inhibitors protrude from the substrate envelope, as these changes selectively weaken inhibitor binding without compromising the binding of substrates. These findings suggest a general model for predicting the susceptibility of protease inhibitors to resistance: drugs designed to fit within the substrate envelope will be less susceptible to resistance, as mutations affecting inhibitor binding would simultaneously interfere with the recognition of viral substrates.
- Department of Biochemistry and Molecular Pharmacology, University of Massachusetts Medical School, Worcester, MA 01605, USA.
Organizational Affiliation: