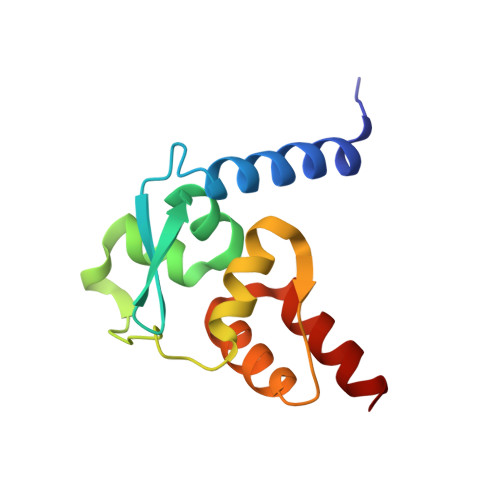
Insights into Strand Exchange in BTB Domain Dimers from the Crystal Structures of FAZF and Miz1.
Stogios, P.J., Cuesta-Seijo, J.A., Chen, L., Pomroy, N.C., Prive, G.G.(2010) J Mol Biology 400: 983-997
- PubMed: 20493880 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2010.05.028
- Primary Citation Related Structures:
3M52, 3M5B - PubMed Abstract:
The BTB domain is a widely distributed protein-protein interaction motif that is often found at the N-terminus of zinc finger transcription factors. Previous crystal structures of BTB domains have revealed tightly interwound homodimers, with the N-terminus from one chain forming a two-stranded anti-parallel beta-sheet with a strand from the other chain. We have solved the crystal structures of the BTB domains from Fanconi anemia zinc finger (FAZF) and Miz1 (Myc-interacting zinc finger 1) to resolutions of 2.0 A and 2.6 A, respectively. Unlike previous examples of BTB domain structures, the FAZF BTB domain is a nonswapped dimer, with each N-terminal beta-strand associated with its own chain. As a result, the dimerization interface in the FAZF BTB domain is about half as large as in the domain-swapped dimers. The Miz1 BTB domain resembles a typical swapped BTB dimer, although it has a shorter N-terminus that is not able to form the interchain sheet. Using cysteine cross-linking, we confirmed that the promyelocytic leukemia zinc finger (PLZF) BTB dimer is strand exchanged in solution, while the FAZF BTB dimer is not. A phylogenic tree of the BTB fold based on both sequence and structural features shows that the common ancestor of the BTB domain in BTB-ZF (bric à brac, tramtrack, broad-complex zinc finger) proteins was a domain-swapped dimer. The differences in the N-termini seen in the FAZF and Miz1 BTB domains appear to be more recent developments in the structural evolution of the domain.
- Department of Medical Biophysics, University of Toronto, Toronto, Ontario, Canada.
Organizational Affiliation: