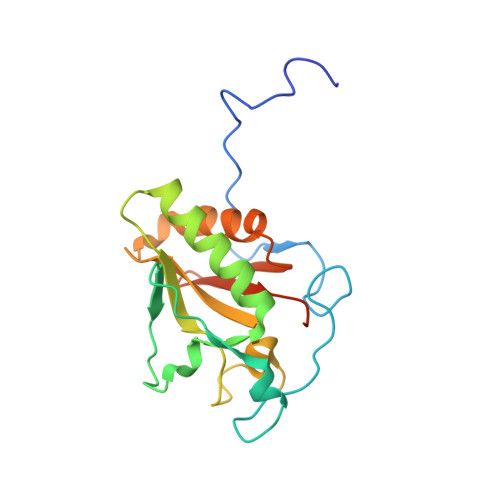
Mapping sonic hedgehog-receptor interactions by steric interference.
Pepinsky, R.B., Rayhorn, P., Day, E.S., Dergay, A., Williams, K.P., Galdes, A., Taylor, F.R., Boriack-Sjodin, P.A., Garber, E.A.(2000) J Biological Chem 275: 10995-11001
- PubMed: 10753901 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.275.15.10995
- Primary Citation Related Structures:
3M1N - PubMed Abstract:
We have defined regions in the Sonic hedgehog (Shh) molecule that are important for Patched (Ptc) receptor binding by targeting selected surface amino acid residues with probes of diverse sizes and shapes and assessing the effects of these modifications on function. Eleven amino acid residues that surround the surface of the protein were chosen for these studies and mutated to cysteine residues. These cysteines were then selectively modified with thiol-specific probes, and the modified proteins were tested for hedgehog receptor binding activity and their ability to induce differentiation of C3H10T1/2 cells into osteoblasts. Based on these analyses, approximately one-third of the Shh surface can be modified without effect on function regardless of the size of the attachment. These sites are located near to where the C terminus protrudes from the surface of the protein. All other sites were sensitive to modification, indicating that the interaction of Shh with its primary receptor Ptc is mediated over a large surface of the Shh protein. For sites Asn-50 and Ser-156, function was lost with the smallest of the probes tested, indicating that these residues are in close proximity to the Ptc-binding site. The epitope for the neutralizing mAb 5E1 mapped to a close but distinct region of the structure. The structure-activity data provide a unique view of the interactions between Shh and Ptc that is not readily attainable by conventional mapping strategies.
- Biogen, Inc., Cambridge, Massachusetts 02142, USA.
Organizational Affiliation: