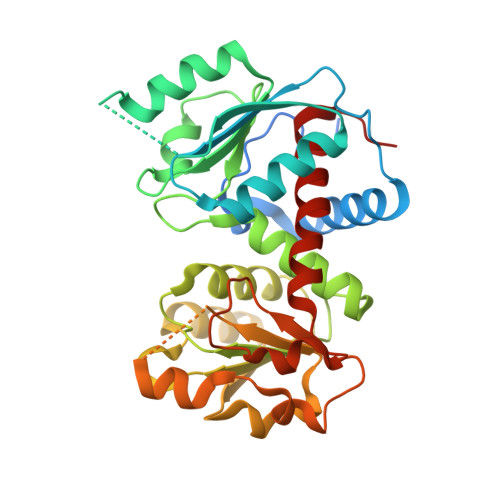
2.00 Angstrom resolution crystal structure of a catalytic subunit of an aspartate carbamoyltransferase (pyrB) from Yersinia pestis CO92
Halavaty, A.S., Minasov, G., Dubrovska, I., Winsor, J., Shuvalova, L., Peterson, S., Anderson, W.F., Center for Structural Genomics of Infectious Diseases (CSGID)To be published.