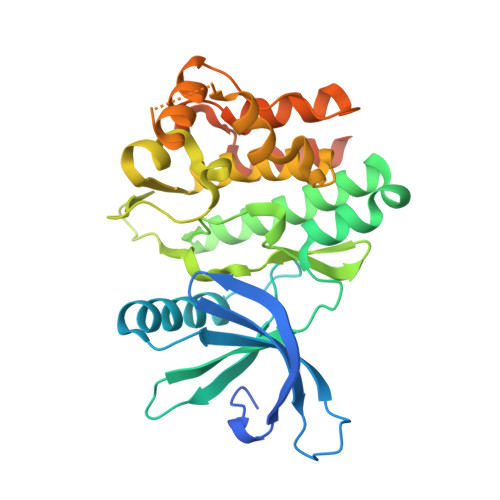
Structural and thermodynamic characterization of the TYK2 and JAK3 kinase domains in complex with CP-690550 and CMP-6.
Chrencik, J.E., Patny, A., Leung, I.K., Korniski, B., Emmons, T.L., Hall, T., Weinberg, R.A., Gormley, J.A., Williams, J.M., Day, J.E., Hirsch, J.L., Kiefer, J.R., Leone, J.W., Fischer, H.D., Sommers, C.D., Huang, H.C., Jacobsen, E.J., Tenbrink, R.E., Tomasselli, A.G., Benson, T.E.(2010) J Mol Biology 400: 413-433
- PubMed: 20478313 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2010.05.020
- Primary Citation Related Structures:
3LXK, 3LXL, 3LXN, 3LXP - PubMed Abstract:
Janus kinases (JAKs) are critical regulators of cytokine pathways and attractive targets of therapeutic value in both inflammatory and myeloproliferative diseases. Although the crystal structures of active JAK1 and JAK2 kinase domains have been reported recently with the clinical compound CP-690550, the structures of both TYK2 and JAK3 with CP-690550 have remained outstanding. Here, we report the crystal structures of TYK2, a first in class structure, and JAK3 in complex with PAN-JAK inhibitors CP-690550 ((3R,4R)-3-[4-methyl-3-[N-methyl-N-(7H-pyrrolo[2,3-d]pyrimidin-4-yl)amino]piperidin-1-yl]-3-oxopropionitrile) and CMP-6 (tetracyclic pyridone 2-t-butyl-9-fluoro-3,6-dihydro-7H-benz[h]-imidaz[4,5-f]isoquinoline-7-one), both of which bind in the ATP-binding cavities of both JAK isozymes in orientations similar to that observed in crystal structures of JAK1 and JAK2. Additionally, a complete thermodynamic characterization of JAK/CP-690550 complex formation was completed by isothermal titration calorimetry, indicating the critical role of the nitrile group from the CP-690550 compound. Finally, computational analysis using WaterMap further highlights the critical positioning of the CP-690550 nitrile group in the displacement of an unfavorable water molecule beneath the glycine-rich loop. Taken together, the data emphasize the outstanding properties of the kinome-selective JAK inhibitor CP-690550, as well as the challenges in obtaining JAK isozyme-selective inhibitors due to the overall structural and sequence similarities between the TYK2, JAK1, JAK2 and JAK3 isozymes. Nevertheless, subtle amino acid variations of residues lining the ligand-binding cavity of the JAK enzymes, as well as the global positioning of the glycine-rich loop, might provide the initial clues to obtaining JAK-isozyme selective inhibitors.
- Pfizer Inc. 700 Chesterfield Parkway West, Chesterfield, MO 63017, USA. jill.chrencik@pfizer.com
Organizational Affiliation: