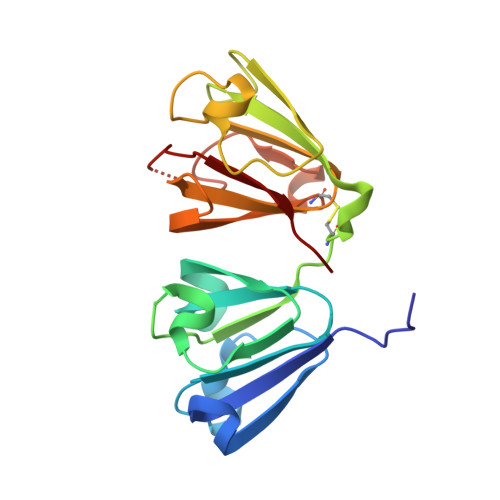
Crystal structure of human Beta-crystallin A4 (CRYBA4)
Chaikuad, A., Shafqat, N., Krojer, T., Yue, W.W., Cocking, R., Vollmar, M., Muniz, J.R.C., Pike, A.C.W., von Delft, F., Arrowsmith, C.H., Weigelt, J., Edwards, A.M., Bountra, C., Oppermann, U.To be published.