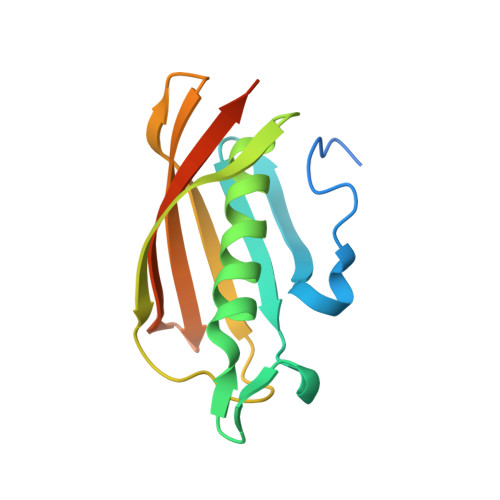
Crystal structure and functional insight of HP0420-homolog from Helicobacter felis
Piao, S., Jin, X.L., Yun, B.-Y., Kim, N., Cho, H.-S., Fukuda, M., Lee, H., Ha, N.-C.(2010) Biochem Biophys Res Commun 394: 940-946
- PubMed: 20302842 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.bbrc.2010.03.087
- Primary Citation Related Structures:
3LW3, 3LWG - PubMed Abstract:
Helicobacter pylori infect more than half of the world's population and are considered a cause of peptic ulcer disease and gastric cancer. Recently, hypothetical gene HP0421 was identified in H. pylori as a cholesterol alpha-glucosyltransferase, which is required to synthesize cholesteryl glucosides, essential cell wall components of the bacteria. In the same gene-cluster, HP0420 was co-identified, whose function remains unknown. Here we report the crystal structure of HP0420-homolog of H. felis (HF0420) to gain insight into the function of HP0420. The crystal structure, combined with size-exclusion chromatography, reveals that HF0420 adopts a homodimeric hot-dog fold. The crystal structure suggests that HF0420 has enzymatic activity that involves a conserved histidine residue at the end of the central alpha-helix. Subsequent biochemical studies provide clues to the function of HP0420 and HF0420.
- College of Pharmacy and Research Institute for Drug Development, Pusan National University, Jangjeon-dong, Geumjeong-gu, Busan 609-735, Republic of Korea.
Organizational Affiliation: