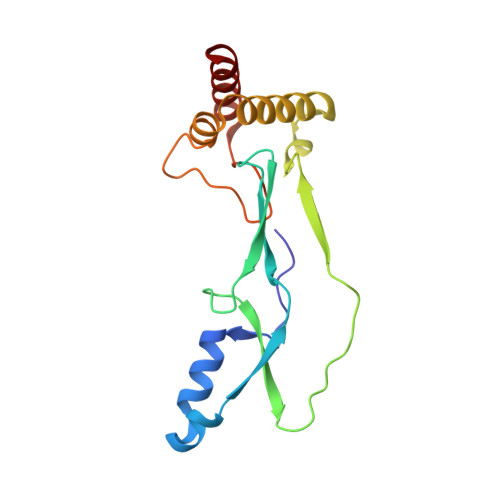
Context-dependent remodeling of structure in two large protein fragments.
Schellenberg, M.J., Ritchie, D.B., Wu, T., Markin, C.J., Spyracopoulos, L., MacMillan, A.M.(2010) J Mol Biology 402: 720-730
- PubMed: 20713060 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2010.08.020
- Primary Citation Related Structures:
3LRU - PubMed Abstract:
Protein folding involves the formation of secondary structural elements from the primary sequence and their association with tertiary assemblies. The relation of this primary sequence to a specific folded protein structure remains a central question in structural biology. An increasing body of evidence suggests that variations in homologous sequence ranging from point mutations to substantial insertions or deletions can yield stable proteins with markedly different folds. Here we report the structural characterization of domain IV (D4) and ΔD4 (polypeptides with 222 and 160 amino acids, respectively) that differ by virtue of an N-terminal deletion of 62 amino acids (28% of the overall D4 sequence). The high-resolution crystal structures of the monomeric D4 and the dimeric ΔD4 reveal substantially different folds despite an overall conservation of secondary structure. These structures show that the formation of tertiary structures, even in extended polypeptide sequences, can be highly context dependent, and they serve as a model for structural plasticity in protein isoforms.
- Department of Biochemistry, School of Molecular and Systems Medicine, University of Alberta, Edmonton, Alberta, Canada.
Organizational Affiliation: