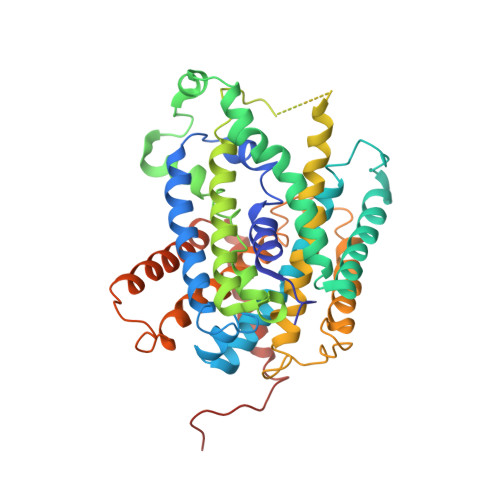
Structure and mechanism of an amino acid antiporter
Gao, X., Lu, F., Zhou, L., Dang, S., Sun, L., Li, X., Wang, J., Shi, Y.(2009) Science 324: 1565-1568
- PubMed: 19478139 Search on PubMed
- DOI: https://doi.org/10.1126/science.1173654
- Primary Citation Related Structures:
3LRB, 3LRC - PubMed Abstract:
Virulent enteric pathogens such as Escherichia coli strain O157:H7 rely on acid-resistance (AR) systems to survive the acidic environment in the stomach. A major component of AR is an arginine-dependent arginine:agmatine antiporter that expels intracellular protons. Here, we report the crystal structure of AdiC, the arginine:agmatine antiporter from E. coli O157:H7 and a member of the amino acid/polyamine/organocation (APC) superfamily of transporters at 3.6 A resolution. The overall fold is similar to that of several Na+-coupled symporters. AdiC contains 12 transmembrane segments, forms a homodimer, and exists in an outward-facing, open conformation in the crystals. A conserved, acidic pocket opens to the periplasm. Structural and biochemical analysis reveals the essential ligand-binding residues, defines the transport route, and suggests a conserved mechanism for the antiporter activity.
- State Key Laboratory of Bio-membrane and Membrane Biotechnology, Tsinghua University, Beijing 100084, China.
Organizational Affiliation: