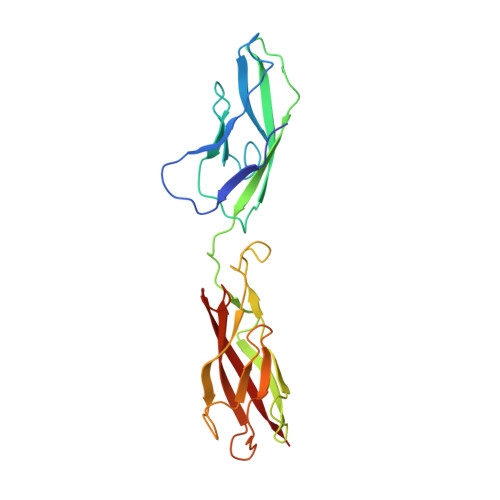
Two-step adhesive binding by classical cadherins.
Harrison, O.J., Bahna, F., Katsamba, P.S., Jin, X., Brasch, J., Vendome, J., Ahlsen, G., Carroll, K.J., Price, S.R., Honig, B., Shapiro, L.(2010) Nat Struct Mol Biol 17: 348-357
- PubMed: 20190754 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nsmb.1784
- Primary Citation Related Structures:
3LND, 3LNE, 3LNF, 3LNG, 3LNH, 3LNI - PubMed Abstract:
Crystal structures of classical cadherins have revealed two dimeric configurations. In the first, N-terminal beta-strands of EC1 domains 'swap' between partner molecules. The second configuration (the 'X dimer'), also observed for T-cadherin, is mediated by residues near the EC1-EC2 calcium binding sites, and N-terminal beta-strands of partner EC1 domains, though held adjacent, do not swap. Here we show that strand-swapping mutants of type I and II classical cadherins form X dimers. Mutant cadherins impaired for X-dimer formation show no binding in short-time frame surface plasmon resonance assays, but in long-time frame experiments, they have homophilic binding affinities close to that of wild type. Further experiments show that exchange between monomers and dimers is slowed in these mutants. These results reconcile apparently disparate results from prior structural studies and suggest that X dimers are binding intermediates that facilitate the formation of strand-swapped dimers.
- Department of Biochemistry and Molecular Biophysics, Columbia University, New York, New York, USA.
Organizational Affiliation: