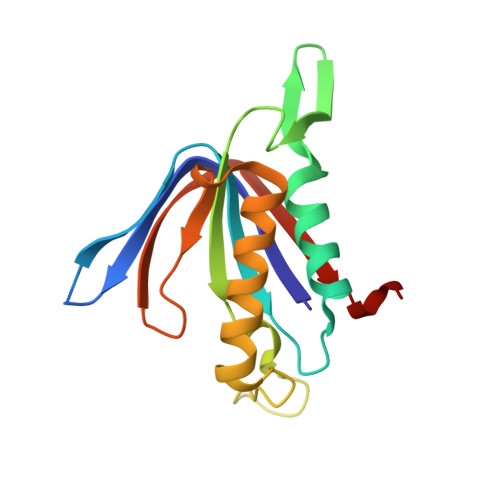
Structure of D-tyrosyl-tRNATyr deacylase using home-source Cu Kalpha and moderate-quality iodide-SAD data: structural polymorphism and HEPES-bound enzyme states
Yogavel, M., Khan, S., Bhatt, T.K., Sharma, A.(2010) Acta Crystallogr D Biol Crystallogr 66: 584-592
- PubMed: 20445234 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444910006062
- Primary Citation Related Structures:
3LMT, 3LMU, 3LMV - PubMed Abstract:
D-Tyrosyl-tRNA(Tyr) deacylase (DTD) is an editing enzyme that removes D-amino acids from mischarged tRNAs. The crystal structure of Plasmodium falciparum DTD (PfDTD) was determined using the iodide-SAD phasing method. Iodide-derivatized PfDTD crystals were obtained using the quick cryo-soaking procedure in which native crystals were soaked for a short period of 10-30 s in cryoprotectant solution containing 0.2-1 M NaI. Iodide-SAD data sets were collected to 3.3 and 2.74 A resolution from PfDTD crystals that belonged to two different space groups, P4(3) and P1, using an in-house X-ray copper-anode source. This is the first report to detail structure solution using low iodide anomalous signal, modest resolution and redundancy and average solvent content for SAD phasing of 984 and 1312 amino acids in the triclinic P1 and tetragonal P4(3) space groups, respectively. A total of 85% and 56% of the residues were automatically built into the iodide-phased electron-density maps using PHENIX AutoBuild. The structure of HEPES-bound PfDTD was subsequently determined by molecular replacement and refined to 2.83 A resolution. The crystals obtained from various batches of crystallization trials of PfDTD exhibited polymorphism in terms of belonging to different crystal forms and space groups. Even within a given crystal system the unit-cell parameters showed high non-isomorphism. These packing variations were exploited in order to conduct a systematic study of conformational changes in PfDTD. It is shown that the disposition of a ten-residue insertion loop affects packing within the PfDTD crystals and seems to determine the non-isomorphism in unit-cell parameters. By tracking the changes in PfDTD unit cells, it was possible to map conformational differences within PfDTD that may be of significance for enzyme activity.
- Structural and Computational Biology Group, International Centre for Genetic Engineering and Biotechnology, Aruna Asaf Ali Road, New Delhi 110 067, India.
Organizational Affiliation: