Inhibitors of hepatitis C virus polymerase: synthesis and characterization of novel 2-oxy-6-fluoro-N-((S)-1-hydroxy-3-phenylpropan-2-yl)-benzamides.
Cheng, C.C., Shipps, G.W., Yang, Z., Kawahata, N., Lesburg, C.A., Duca, J.S., Bandouveres, J., Bracken, J.D., Jiang, C.K., Agrawal, S., Ferrari, E., Huang, H.C.(2010) Bioorg Med Chem Lett 20: 2119-2124
- PubMed: 20219368 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmcl.2010.02.054
- Primary Citation Related Structures:
3LKH - PubMed Abstract:
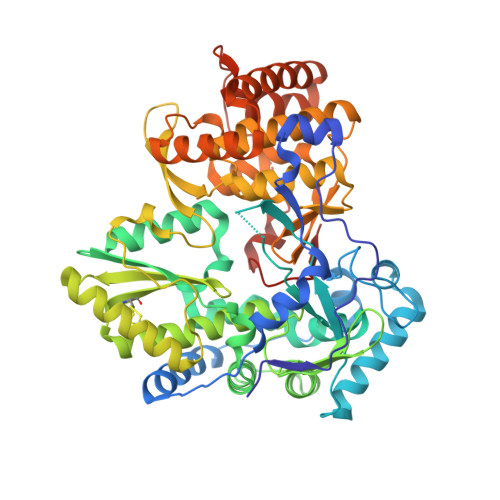
SAR exploration from an initial hit, (S)-N-(2-cyclohexenylethyl)-2-fluoro-6-(2-(1-hydroxy-3-phenylpropan-2-ylamino)-2-oxoethoxy)benzamide (1), identified using our proprietary automated ligand identification system (ALIS),(1) has led to a novel series of selective hepatitis C virus (HCV) NS5B polymerase inhibitors with improved in vitro potency as exemplified by (S)-2-fluoro-6-(2-(1-hydroxy-3-phenylpropan-2-ylamino)-2-oxoethoxy)-N-isopentyl-N-methylbenzamidecarboxamide (41) (IC(50)=0.5 microM). The crystal structure of an analogue (44) was solved and provided rationalization of the SAR of this series, which binds in a distinct manner in the palm domain of NS5B, consistent with biochemical analysis using enzyme mutant variants. These data warrant further lead optimization efforts on this novel series of non-nucleoside inhibitors targeting the HCV polymerase.
- Department of Lead Discovery, Merck Research Laboratories, 320 Bent Street, Cambridge, MA 02141, United States. cccheng@alum.mit.edu
Organizational Affiliation: