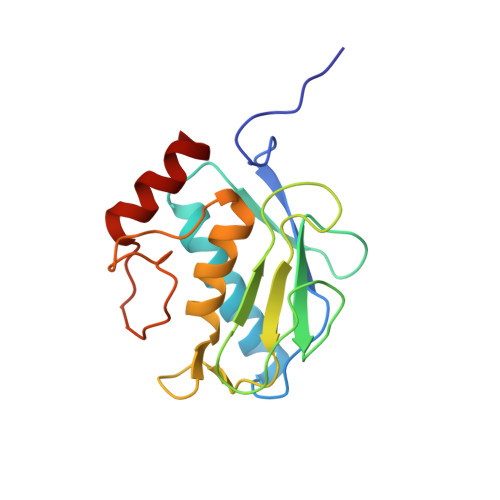
Structure analysis reveals the flexibility of the ADAMTS-5 active site.
Shieh, H.S., Tomasselli, A.G., Mathis, K.J., Schnute, M.E., Woodard, S.S., Caspers, N., Williams, J.M., Kiefer, J.R., Munie, G., Wittwer, A., Malfait, A.M., Tortorella, M.D.(2011) Protein Sci 20: 735-744
- PubMed: 21370305 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.606
- Primary Citation Related Structures:
3LJT, 3LJZ - PubMed Abstract:
A ((1S,2R)-2-hydroxy-2,3-dihydro-1H-inden-1-yl) succinamide derivative (here referred to as Compound 12) shows significant activity toward many matrix metalloproteinases (MMPs), including MMP-2, MMP-8, MMP-9, and MMP-13. Modeling studies had predicted that this compound would not bind to ADAMTS-5 (a disintegrin and metalloproteinase with thrombospondin motifs-5) due to its shallow S1' pocket. However, inhibition analysis revealed it to be a nanomolar inhibitor of both ADAMTS-4 and -5. The observed inconsistency was explained by analysis of crystallographic structures, which showed that Compound 12 in complex with the catalytic domain of ADAMTS-5 (cataTS5) exhibits an unusual conformation in the S1' pocket of the protein. This first demonstration that cataTS5 can undergo an induced conformational change in its active site pocket by a molecule like Compound 12 should enable the design of new aggrecanase inhibitors with better potency and selectivity profiles.
- Pfizer Global Research and Development, St. Louis, Missouri 63017, USA. shiehouse@sbcglobal.net
Organizational Affiliation: