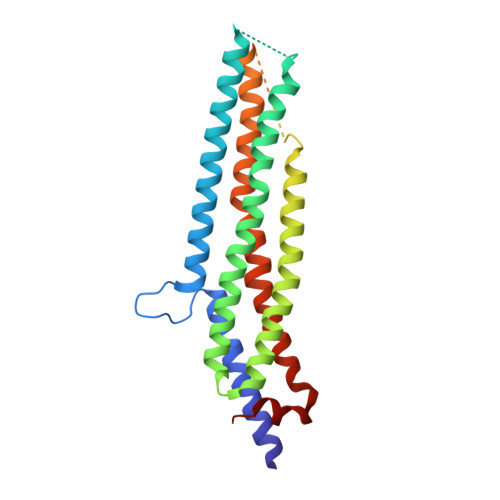
Structural basis of oligomerization in the stalk region of dynamin-like MxA.
Gao, S., von der Malsburg, A., Paeschke, S., Behlke, J., Haller, O., Kochs, G., Daumke, O.(2010) Nature 465: 502-506
- PubMed: 20428112 Search on PubMed
- DOI: https://doi.org/10.1038/nature08972
- Primary Citation Related Structures:
3LJB - PubMed Abstract:
The interferon-inducible dynamin-like myxovirus resistance protein 1 (MxA; also called MX1) GTPase is a key mediator of cell-autonomous innate immunity against pathogens such as influenza viruses. MxA partially localizes to COPI-positive membranes of the smooth endoplasmic reticulum-Golgi intermediate compartment. At the point of infection, it redistributes to sites of viral replication and promotes missorting of essential viral constituents. It has been proposed that the middle domain and the GTPase effector domain of dynamin-like GTPases constitute a stalk that mediates oligomerization and transmits conformational changes from the G domain to the target structure; however, the molecular architecture of this stalk has remained elusive. Here we report the crystal structure of the stalk of human MxA, which folds into a four-helical bundle. This structure tightly oligomerizes in the crystal in a criss-cross pattern involving three distinct interfaces and one loop. Mutations in each of these interaction sites interfere with native assembly, oligomerization, membrane binding and antiviral activity of MxA. On the basis of these results, we propose a structural model for dynamin oligomerization and stimulated GTP hydrolysis that is consistent with previous structural predictions and has functional implications for all members of the dynamin family.
- Max-Delbrück-Centrum for Molecular Medicine, Crystallography, Robert-Rössle-Strasse 10, 13125 Berlin, Germany.
Organizational Affiliation: