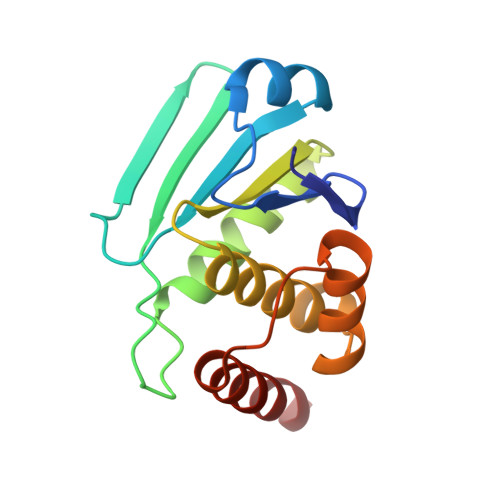
Exploring binding sites other than the catalytic core in the crystal structure of the catalytic domain of MKP-4
Jeong, D.G., Yoon, T.S., Jung, S.-K., Park, B.C., Park, H.S., Ryu, S.E., Kim, S.J.(2011) Acta Crystallogr D Biol Crystallogr 67: 25-31
- PubMed: 21206059 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444910042381
- Primary Citation Related Structures:
3LJ8 - PubMed Abstract:
Map kinase phosphatase 4 (MKP-4), which has been implicated in signalling pathways that negatively regulate glucose uptake, belongs to the dual-specificity phosphatase (DUSP) family. An inherent property of MKPs is an ability to undergo structural rearrangement, transitioning from a partially active to a fully active conformation. Here, a 2.7 Å resolution crystal structure of the catalytic domain of MKP-4 (MKP-4C) is presented. It was determined that the MKP-4C structure seriously deviates from canonical conformations of DUSPs and this characteristic feature results in significant gaps between the catalytic core and several surrounding loops which are unique compared with other MKP counterparts that adopt an active conformation. Using virtual library screening, it was found that inhibitors bind to MKP-4C with high affinity near these gaps. Inhibitors that target other binding sites instead of the active site can be utilized to prevent transition to a fully active conformation. Compounds that are able to make contacts with these sites in MKP-4 would not only provide a beneficial increase in affinity but may also permit greater specificity relative to other protein tyrosine phosphatases.
- Medical Proteomics Research Center, Korea Research Institute of Bioscience and Biotechnology, Yuseong-Gu, Daejeon, Republic of Korea.
Organizational Affiliation: