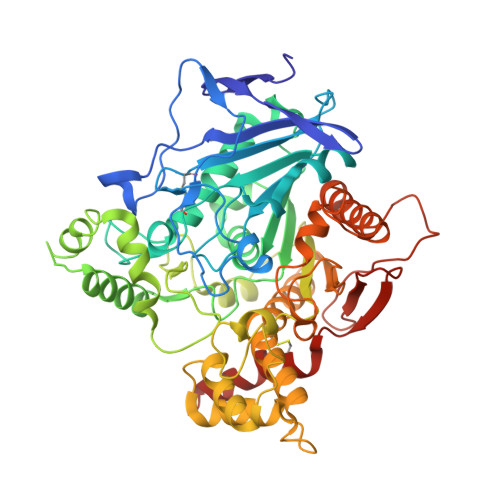
Acetylcholinesterase: From 3D structure to function.
Dvir, H., Silman, I., Harel, M., Rosenberry, T.L., Sussman, J.L.(2010) Chem Biol Interact 187: 10-22
- PubMed: 20138030
- DOI: https://doi.org/10.1016/j.cbi.2010.01.042
- Primary Citation of Related Structures:
3LII - PubMed Abstract:
By rapid hydrolysis of the neurotransmitter, acetylcholine, acetylcholinesterase terminates neurotransmission at cholinergic synapses. Acetylcholinesterase is a very fast enzyme, functioning at a rate approaching that of a diffusion-controlled reaction. The powerful toxicity of organophosphate poisons is attributed primarily to their potent inhibition of acetylcholinesterase. Acetylcholinesterase inhibitors are utilized in the treatment of various neurological disorders, and are the principal drugs approved thus far by the FDA for management of Alzheimer's disease. Many organophosphates and carbamates serve as potent insecticides, by selectively inhibiting insect acetylcholinesterase. The determination of the crystal structure of Torpedo californica acetylcholinesterase permitted visualization, for the first time, at atomic resolution, of a binding pocket for acetylcholine. It also allowed identification of the active site of acetylcholinesterase, which, unexpectedly, is located at the bottom of a deep gorge lined largely by aromatic residues. The crystal structure of recombinant human acetylcholinesterase in its apo-state is similar in its overall features to that of the Torpedo enzyme; however, the unique crystal packing reveals a novel peptide sequence which blocks access to the active-site gorge.
- Department of Structural Biology, Weizmann Institute of Science, Rehovot 76100, Israel.
Organizational Affiliation: