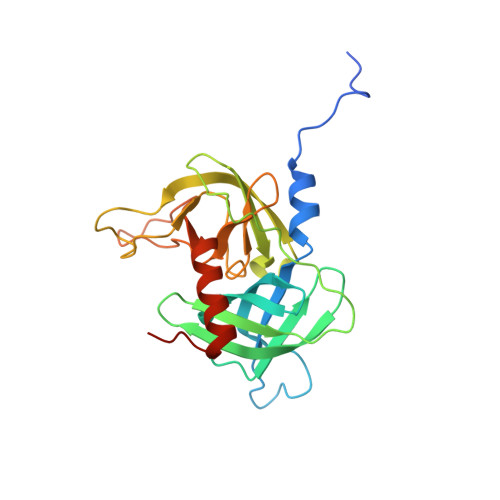
Allostery is an intrinsic property of the protease domain of DegS: implications for enzyme function and evolution.
Sohn, J., Grant, R.A., Sauer, R.T.(2010) J Biological Chem 285: 34039-34047
- PubMed: 20739286 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M110.135541
- Primary Citation Related Structures:
3LGI, 3LGT, 3LGU, 3LGV, 3LGW, 3LGY, 3LH1, 3LH3 - PubMed Abstract:
DegS is a periplasmic Escherichia coli protease, which functions as a trimer to catalyze the initial rate-limiting step in a proteolytic cascade that ultimately activates transcription of stress response genes in the cytoplasm. Each DegS subunit consists of a protease domain and a PDZ domain. During protein folding stress, DegS is allosterically activated by peptides exposed in misfolded outer membrane porins, which bind to the PDZ domain and stabilize the active protease. It is not known whether allostery is conferred by the PDZ domains or is an intrinsic feature of the trimeric protease domain. Here, we demonstrate that free DegS(ΔPDZ) equilibrates between active and inactive trimers with the latter species predominating. Substrate binding stabilizes active DegS(ΔPDZ) in a positively cooperative fashion. Mutations can also stabilize active DegS(ΔPDZ) and produce an enzyme that displays hyperbolic kinetics and degrades substrate with a maximal velocity within error of that for fully activated, intact DegS. Crystal structures of multiple DegS(ΔPDZ) variants, in functional and non-functional conformations, support a two-state model in which allosteric switching is mediated by changes in specific elements of tertiary structure in the context of an invariant trimeric base. Overall, our results indicate that protein substrates must bind sufficiently tightly and specifically to the functional conformation of DegS(ΔPDZ) to assist their own degradation. Thus, substrate binding alone may have regulated the activities of ancestral DegS trimers with subsequent fusion of the protease domain to a PDZ domain, resulting in ligand-mediated regulation.
- Department of Biology, Massachusetts Institute of Technology, Cambridge, Massachusetts 02139, USA.
Organizational Affiliation: