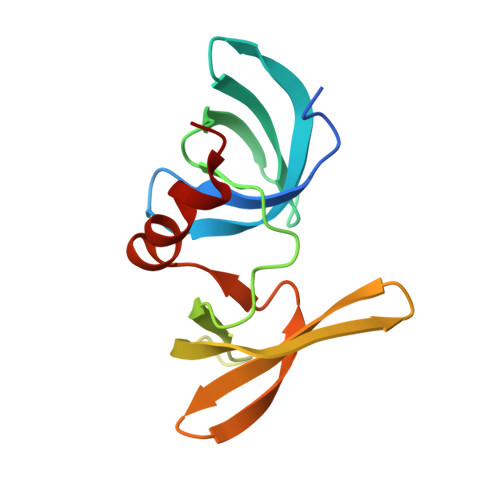
Structural insight into p53 recognition by the 53BP1 tandem Tudor domain.
Roy, S., Musselman, C.A., Kachirskaia, I., Hayashi, R., Glass, K.C., Nix, J.C., Gozani, O., Appella, E., Kutateladze, T.G.(2010) J Mol Biology 398: 489-496
- PubMed: 20307547 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2010.03.024
- Primary Citation Related Structures:
3LGF, 3LGL, 3LH0 - PubMed Abstract:
The tumor suppressor p53 and the DNA repair factor 53BP1 (p53 binding protein 1) regulate gene transcription and responses to genotoxic stresses. Upon DNA damage, p53 undergoes dimethylation at Lys382 (p53K382me2), and this posttranslational modification is recognized by 53BP1. The molecular mechanism of nonhistone methyl-lysine mark recognition remains unknown. Here we report a 1. 6-A-resolution crystal structure of the tandem Tudor domain of human 53BP1 bound to a p53K382me2 peptide. In the complex, dimethylated Lys382 is restrained by a set of hydrophobic and cation-pi interactions in a cage formed by four aromatic residues and an aspartate of 53BP1. The signature HKKme2 motif of p53, which defines specificity, is identified through a combination of NMR resonance perturbations, mutagenesis, measurements of binding affinities and docking simulations, and analysis of the crystal structures of 53BP1 bound to p53 peptides containing other dimethyl-lysine marks, p53K370me2 (p53 dimethylated at Lys370) and p53K372me2 (p53 dimethylated at Lys372). Binding of the 53BP1 Tudor domain to p53K382me2 may facilitate p53 accumulation at DNA damage sites and promote DNA repair as suggested by chromatin immunoprecipitation and DNA repair assays. Together, our data detail the molecular mechanism of p53-53BP1 association and provide the basis for deciphering the role of this interaction in the regulation of p53 and 53BP1 functions.
- Department of Pharmacology, University of Colorado Denver School of Medicine, 12801 East 17th Avenue, Aurora, CO 80045-0511, USA.
Organizational Affiliation: