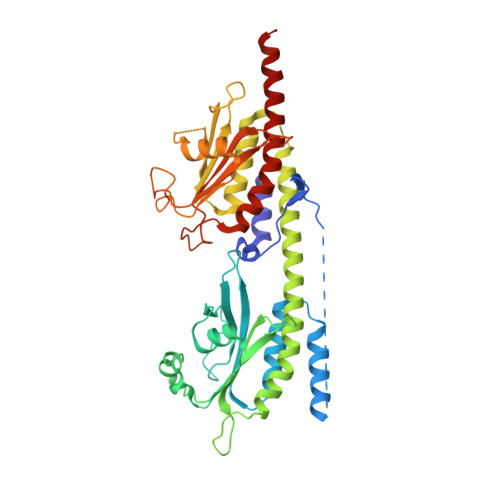
Conformation changes, N-terminal involvement, and cGMP signal relay in the phosphodiesterase-5 GAF domain.
Wang, H., Robinson, H., Ke, H.(2010) J Biological Chem 285: 38149-38156
- PubMed: 20861010 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M110.141614
- Primary Citation Related Structures:
3LFV, 3MF0 - PubMed Abstract:
The activity of phosphodiesterase-5 (PDE5) is specific for cGMP and is regulated by cGMP binding to GAF-A in its regulatory domain. To better understand the regulatory mechanism, x-ray crystallographic and biochemical studies were performed on constructs of human PDE5A1 containing the N-terminal phosphorylation segment, GAF-A, and GAF-B. Superposition of this unliganded GAF-A with the previously reported NMR structure of cGMP-bound PDE5 revealed dramatic conformational differences and suggested that helix H4 and strand B3 probably serve as two lids to gate the cGMP-binding pocket in GAF-A. The structure also identified an interfacial region among GAF-A, GAF-B, and the N-terminal loop, which may serve as a relay of the cGMP signal from GAF-A to GAF-B. N-terminal loop 98-147 was physically associated with GAF-B domains of the dimer. Biochemical analyses showed an inhibitory effect of this loop on cGMP binding and its involvement in the cGMP-induced conformation changes.
- Department of Biochemistry and Biophysics and Lineberger Comprehensive Cancer Center, University of North Carolina, Chapel Hill, North Carolina 27599-7260, USA.
Organizational Affiliation: