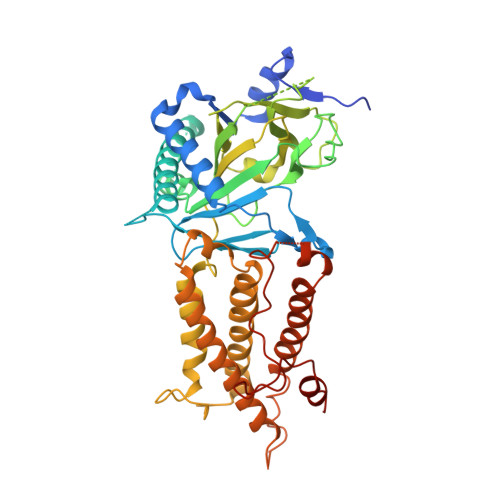
Crystal structure of the FTO protein reveals basis for its substrate specificity
Han, Z., Niu, T., Chang, J., Lei, X., Zhao, M., Wang, Q., Cheng, W., Wang, J., Feng, Y., Chai, J.(2010) Nature 464: 1205-1209
- PubMed: 20376003 Search on PubMed
- DOI: https://doi.org/10.1038/nature08921
- Primary Citation Related Structures:
3LFM - PubMed Abstract:
Recent studies have unequivocally associated the fat mass and obesity-associated (FTO) gene with the risk of obesity. In vitro FTO protein is an AlkB-like DNA/RNA demethylase with a strong preference for 3-methylthymidine (3-meT) in single-stranded DNA or 3-methyluracil (3-meU) in single-stranded RNA. Here we report the crystal structure of FTO in complex with the mononucleotide 3-meT. FTO comprises an amino-terminal AlkB-like domain and a carboxy-terminal domain with a novel fold. Biochemical assays show that these two domains interact with each other, which is required for FTO catalytic activity. In contrast with the structures of other AlkB members, FTO possesses an extra loop covering one side of the conserved jelly-roll motif. Structural comparison shows that this loop selectively competes with the unmethylated strand of the DNA duplex for binding to FTO, suggesting that it has an important role in FTO selection against double-stranded nucleic acids. The ability of FTO to distinguish 3-meT or 3-meU from other nucleotides is conferred by its hydrogen-bonding interaction with the two carbonyl oxygen atoms in 3-meT or 3-meU. Taken together, these results provide a structural basis for understanding FTO substrate-specificity, and serve as a foundation for the rational design of FTO inhibitors.
- National Institute of Biological Sciences, No. 7 Science Park Road, Beijing 102206, China.
Organizational Affiliation: