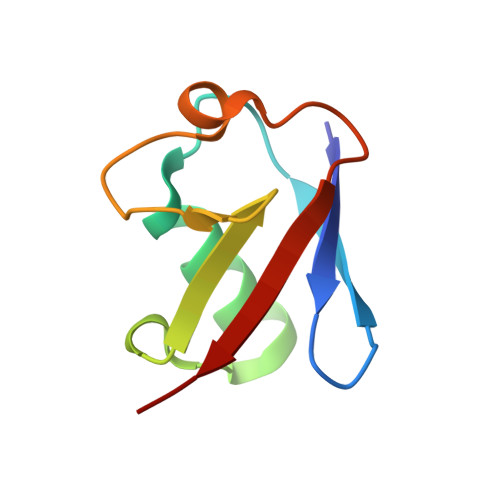
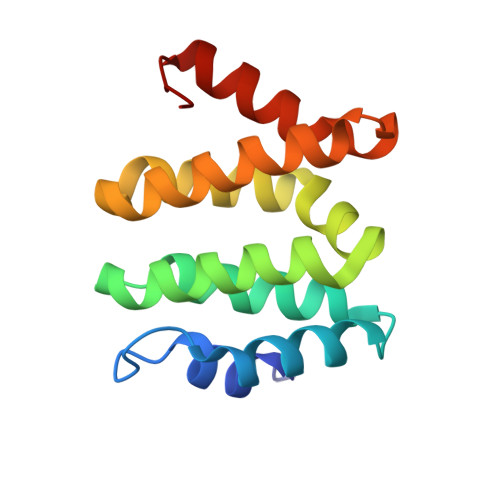
VHS domains of ESCRT-0 cooperate in high-avidity binding to polyubiquitinated cargo.
Ren, X., Hurley, J.H.(2010) EMBO J 29: 1045-1054
- PubMed: 20150893 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/emboj.2010.6
- Primary Citation Related Structures:
3LDZ - PubMed Abstract:
VHS (Vps27, Hrs, and STAM) domains occur in ESCRT-0 subunits Hrs and STAM, GGA adapters, and other trafficking proteins. The structure of the STAM VHS domain-ubiquitin complex was solved at 2.6 A resolution, revealing that determinants for ubiquitin recognition are conserved in nearly all VHS domains. VHS domains from all classes of VHS-domain containing proteins in yeast and humans, including both subunits of ESCRT-0, bound ubiquitin in vitro. ESCRTs have been implicated in the sorting of Lys63-linked polyubiquitinated cargo. Intact human ESCRT-0 binds Lys63-linked tetraubiquitin 50-fold more tightly than monoubiquitin, though only 2-fold more tightly than Lys48-linked tetraubiquitin. The gain in affinity is attributed to the cooperation of flexibly connected VHS and UIM motifs of ESCRT-0 in avid binding to the polyubiquitin chain. Mutational analysis of all the five ubiquitin-binding sites in yeast ESCRT-0 shows that cooperation between them is required for the sorting of the Lys63-linked polyubiquitinated cargo Cps1 to the vacuole.
- Laboratory of Molecular Biology, National Institute of Diabetes and Digestive and Kidney Diseases, National Institutes of Health, US Department of Health and Human Services, Bethesda, MD, USA.
Organizational Affiliation: