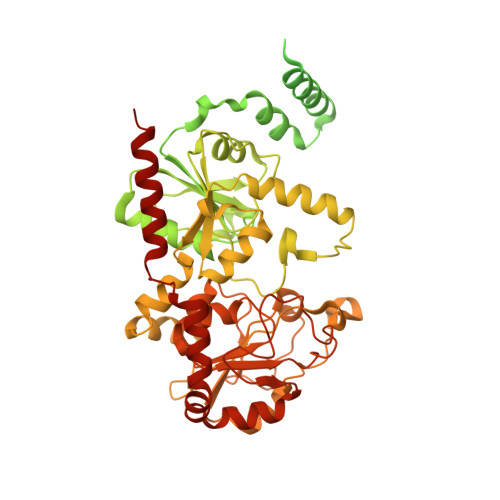
Structure of the bacterial teichoic acid polymerase TagF provides insights into membrane association and catalysis.
Lovering, A.L., Lin, L.Y., Sewell, E.W., Spreter, T., Brown, E.D., Strynadka, N.C.(2010) Nat Struct Mol Biol 17: 582-589
- PubMed: 20400947 Search on PubMed
- DOI: https://doi.org/10.1038/nsmb.1819
- Primary Citation Related Structures:
3L7I, 3L7J, 3L7K, 3L7L, 3L7M - PubMed Abstract:
Teichoic acid polymers are composed of polyol-phosphate units and form a major component of Gram-positive bacterial cell walls. These anionic compounds perform a multitude of important roles in bacteria and are synthesized by monotopic membrane proteins of the TagF polymerase family. We have determined the structure of Staphylococcus epidermidis TagF to 2.7-A resolution from a construct that includes both the membrane-targeting region and the glycerol-phosphate polymerase domains. TagF possesses a helical region for interaction with the lipid bilayer, placing the active site at a suitable distance for access to the membrane-bound substrate. Characterization of active-site residue variants and analysis of a CDP-glycerol substrate complex suggest a mechanism for polymer synthesis. With the importance of teichoic acid in Gram-positive physiology, this elucidation of the molecular details of TagF function provides a critical new target in the development of novel anti-infectives.
- Department of Biochemistry and Molecular Biology, Vancouver, Canada.
Organizational Affiliation: