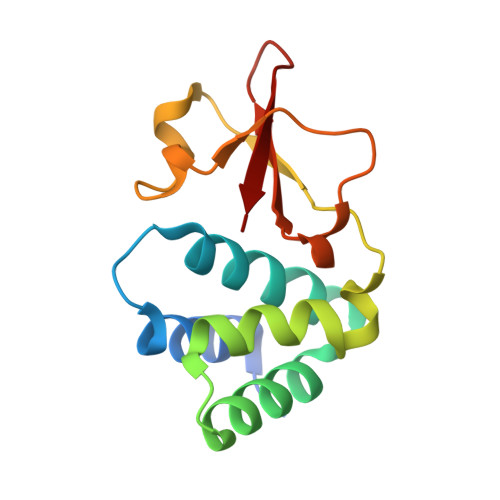
Structural basis for dsRNA recognition and interferon antagonism by Ebola VP35.
Leung, D.W., Prins, K.C., Borek, D.M., Farahbakhsh, M., Tufariello, J.M., Ramanan, P., Nix, J.C., Helgeson, L.A., Otwinowski, Z., Honzatko, R.B., Basler, C.F., Amarasinghe, G.K.(2010) Nat Struct Mol Biol 17: 165-172
- PubMed: 20081868 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nsmb.1765
- Primary Citation Related Structures:
3L25, 3L26, 3L27, 3L28 - PubMed Abstract:
Ebola viral protein 35 (VP35), encoded by the highly pathogenic Ebola virus, facilitates host immune evasion by antagonizing antiviral signaling pathways, including those initiated by RIG-I-like receptors. Here we report the crystal structure of the Ebola VP35 interferon inhibitory domain (IID) bound to short double-stranded RNA (dsRNA), which together with in vivo results reveals how VP35-dsRNA interactions contribute to immune evasion. Conserved basic residues in VP35 IID recognize the dsRNA backbone, whereas the dsRNA blunt ends are 'end-capped' by a pocket of hydrophobic residues that mimic RIG-I-like receptor recognition of blunt-end dsRNA. Residues critical for RNA binding are also important for interferon inhibition in vivo but not for viral polymerase cofactor function of VP35. These results suggest that simultaneous recognition of dsRNA backbone and blunt ends provides a mechanism by which Ebola VP35 antagonizes host dsRNA sensors and immune responses.
- Department of Biochemistry, Biophysics and Molecular Biology, Iowa State University, Ames, Iowa, USA.
Organizational Affiliation: