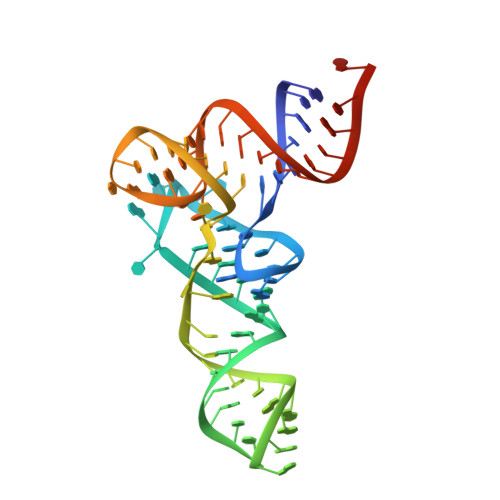
The crystal structure of unmodified tRNAPhe from Escherichia coli
Byrne, R.T., Konevega, A.L., Rodnina, M.V., Antson, A.A.(2010) Nucleic Acids Res 38: 4154-4162
- PubMed: 20203084 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkq133
- Primary Citation Related Structures:
3L0U - PubMed Abstract:
Post-transcriptional nucleoside modifications fine-tune the biophysical and biochemical properties of transfer RNA (tRNA) so that it is optimized for participation in cellular processes. Here we report the crystal structure of unmodified tRNA(Phe) from Escherichia coli at a resolution of 3 A. We show that in the absence of modifications the overall fold of the tRNA is essentially the same as that of mature tRNA. However, there are a number of significant structural differences, such as rearrangements in a triplet base pair and a widened angle between the acceptor and anticodon stems. Contrary to previous observations, the anticodon adopts the same conformation as seen in mature tRNA.
- York Structural Biology Laboratory, Department of Chemistry, University of York, Heslington, North Yorkshire, YO10 5YW, UK.
Organizational Affiliation: