Structural mechanism of host Rab1 activation by the bifunctional Legionella type IV effector SidM/DrrA
Zhu, Y., Hu, L., Zhou, Y., Yao, Q., Liu, L., Shao, F.(2010) Proc Natl Acad Sci U S A 107: 4699-4704
- PubMed: 20176951 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0914231107
- Primary Citation Related Structures:
3L0I, 3L0M - PubMed Abstract:
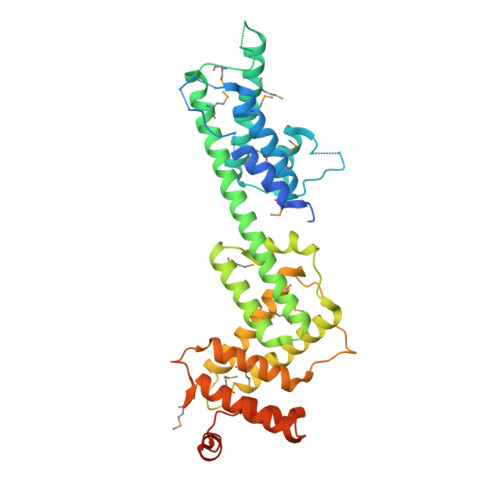
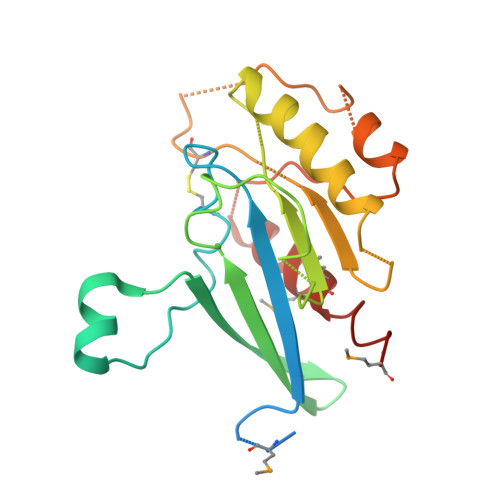
Bacterial pathogens deliver effector proteins with diverse biochemical activities into host cells, thereby modulating various host functions. Legionella pneumophila hijacks host vesicle trafficking to avoid phagosome-lysosome fusion, a mechanism that is dependent on the Legionella Dot/Icm type IV secretion system. SidM/DrrA, a Legionella type IV effector, is important for the interactions of Legionella-containing vacuoles with host endoplasmic reticulum-derived vesicles. SidM is the only known protein that catalyzes both the exchange of GDP for GTP and GDI displacement from small GTPase Rab1. We determined the crystal structures of SidM alone (residues 317-647) and SidM (residues 193-550) in complex with nucleotide-free WT Rab1. The SidM structure contains an N-terminal helical domain with a potential new function, a Rab1-activation domain, and a C-terminal phosphatidylinositol 4-phosphate-binding P4M domain. The Rab1-activation domain has extensive strong interactions mainly with Rab1 switch I and II regions that undergo substantial conformational changes on SidM binding. Mutations of switch-contacting residues in SidM attenuate both the nucleotide exchange and GDI displacement activities. Structural comparisons of Rab1 in the SidM complex with Rab1-GDP and Ypt1-GDP in the GDI complex identify key conformational changes that disrupt the nucleotide and GDI binding of Rab1. Further biochemical and structural analyses reveal a unique mechanism of coupled GDP release and GDI displacement likely triggered by the SidM-induced drastic displacement of switch I of Rab1.
- National Institute of Biological Sciences, Beijing 102206, China.
Organizational Affiliation: