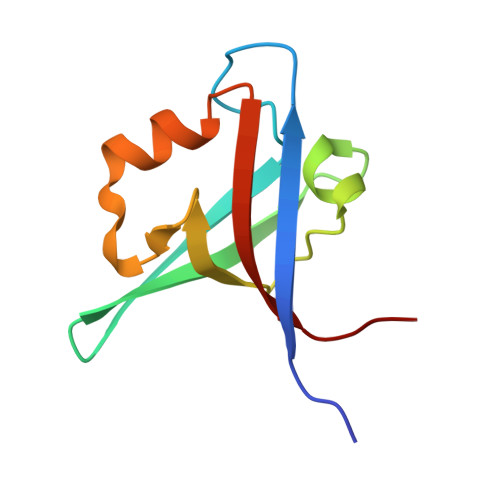
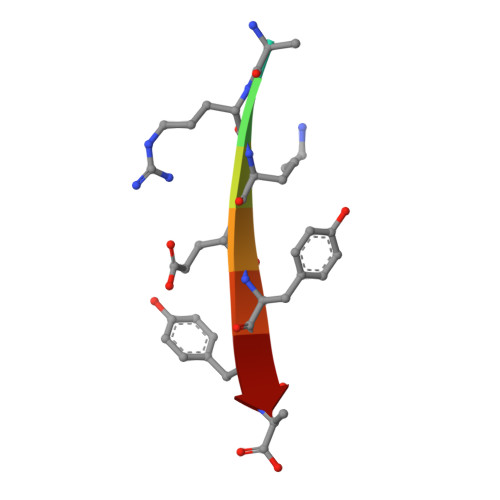
The Tiam1 PDZ domain couples to Syndecan1 and promotes cell-matrix adhesion.
Shepherd, T.R., Klaus, S.M., Liu, X., Ramaswamy, S., DeMali, K.A., Fuentes, E.J.(2010) J Mol Biology 398: 730-746
- PubMed: 20361982 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2010.03.047
- Primary Citation Related Structures:
3KZD, 3KZE - PubMed Abstract:
The T-cell lymphoma invasion and metastasis gene 1 (Tiam1) is a guanine exchange factor (GEF) for the Rho-family GTPase Rac1 that is crucial for the integrity of adherens junctions, tight junctions, and cell-matrix interactions. This GEF contains several protein-protein interaction domains, including a PDZ domain. Earlier studies identified a consensus PDZ-binding motif and a synthetic peptide capable of binding to the Tiam1 PDZ domain, but little is known about its ligand specificity and physiological role in cells. Here, we investigated the structure, specificity, and function of the Tiam1 PDZ domain. We determined the crystal structures of the Tiam1 PDZ domain free and in complex with a "model" peptide, which revealed the structural basis for ligand specificity. Protein database searches using the consensus PDZ-binding motif identified two eukaryotic cell adhesion proteins, Syndecan1 and Caspr4, as potential Tiam1 PDZ domain binding proteins. Equilibrium binding experiments confirmed that C-terminal peptides derived from Syndecan1 and Caspr4 bound the Tiam1 PDZ domain. NMR chemical shift perturbation experiments indicated that the Tiam1 PDZ/Syndecan1 and PDZ/Caspr4 complexes were structurally distinct and identified key residues likely to be responsible for ligand selectivity. Moreover, cell biological analysis established that Syndecan1 is a physiological binding partner of Tiam1 and that the PDZ domain has a function in cell-matrix adhesion and cell migration. Collectively, our data provide insight into the structure, specificity, and function of the Tiam1 PDZ domain. Importantly, our data report on a physiological role for the Tiam1 PDZ domain and establish a novel link between two previously unrelated signal transduction pathways, both of which are implicated in cancer.
- University of Iowa, Department of Biochemistry, Roy J. and Lucille A. Carver College of Medicine, 4-632 Bowen Science Building, Iowa City, IA 52242-1109, USA.
Organizational Affiliation: