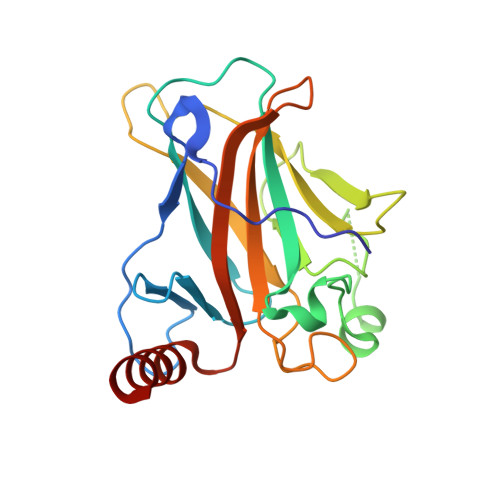
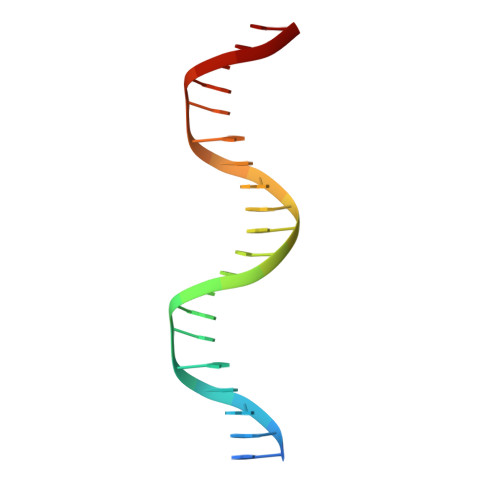
Diversity in DNA recognition by p53 revealed by crystal structures with Hoogsteen base pairs
Kitayner, M., Rozenberg, H., Rohs, R., Suad, O., Rabinovich, D., Honig, B., Shakked, Z.(2010) Nat Struct Mol Biol 17: 423-429
- PubMed: 20364130 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nsmb.1800
- Primary Citation Related Structures:
3IGK, 3IGL, 3KZ8 - PubMed Abstract:
p53 binds as a tetramer to DNA targets consisting of two decameric half-sites separated by a variable spacer. Here we present high-resolution crystal structures of complexes between p53 core-domain tetramers and DNA targets consisting of contiguous half-sites. In contrast to previously reported p53-DNA complexes that show standard Watson-Crick base pairs, the newly reported structures show noncanonical Hoogsteen base-pairing geometry at the central A-T doublet of each half-site. Structural and computational analyses show that the Hoogsteen geometry distinctly modulates the B-DNA helix in terms of local shape and electrostatic potential, which, together with the contiguous DNA configuration, results in enhanced protein-DNA and protein-protein interactions compared to noncontiguous half-sites. Our results suggest a mechanism relating spacer length to protein-DNA binding affinity. Our findings also expand the current understanding of protein-DNA recognition and establish the structural and chemical properties of Hoogsteen base pairs as the basis for a novel mode of sequence readout.
- Department of Structural Biology, Weizmann Institute of Science, Rehovot, Israel.
Organizational Affiliation: