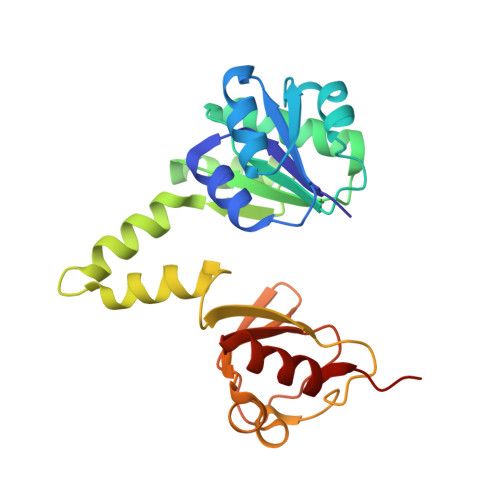
Structure of the MthK RCK in complex with cadmium.
Dvir, H., Valera, E., Choe, S.(2010) J Struct Biol 171: 231-237
- PubMed: 20371380 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jsb.2010.03.020
- Primary Citation Related Structures:
3KXD - PubMed Abstract:
RCK is a cytoplasmic regulatory domain of calcium-gated potassium channels. Binding of Ca(2+) by RCK leads to channel activation through a series of yet unknown conformational changes. Structures of the K(+) channel, MthK, and its cytoplasmic RCK domain revealed two binding sites for Ca(2+) per dimer. We determined the crystal structure of RCK in complex with Cd(2+) at 2.2A resolution. Cd(2+) activates MthK more efficiently, and binds at the same binding sites for Ca(2+) but with reduced coordination number. Two additional binding sites for Cd(2+) are found per dimer; one on the main Rossman-fold lobe, and the other on the small lobe of RCK. Using patch-clamp experiments, we demonstrate that Cd(2+) binding to these novel sites enhances activation by Cd(2+) and not by Ca(2+). The structure reveals a large negatively charged surface patch in the proximity of the Ca(2+)/Cd(2+) binding sites, charge neutralization of which appears to promote the channel open state.
- Structural Biology Laboratory, The Salk Institute for Biological Studies, 10010 North Torrey Pines Road, La Jolla, CA 92037, USA.
Organizational Affiliation: