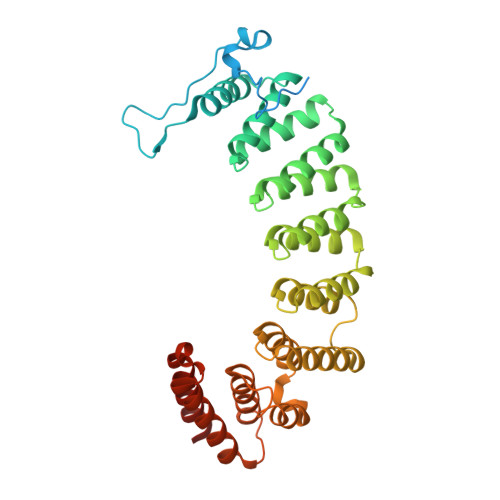
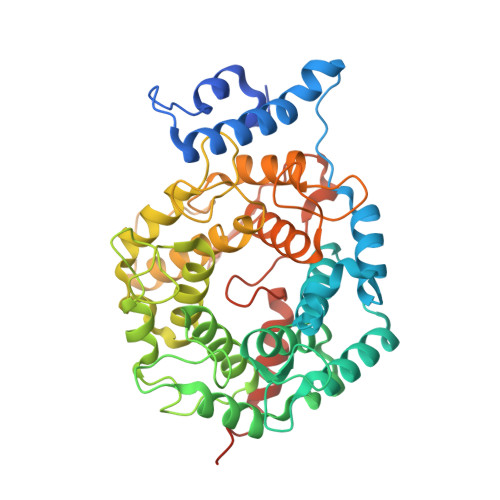
Synthesis, properties, and applications of diazotrifluropropanoyl-containing photoactive analogs of farnesyl diphosphate containing modified linkages for enhanced stability.
Hovlid, M.L., Edelstein, R.L., Henry, O., Ochocki, J., DeGraw, A., Lenevich, S., Talbot, T., Young, V.G., Hruza, A.W., Lopez-Gallego, F., Labello, N.P., Strickland, C.L., Schmidt-Dannert, C., Distefano, M.D.(2010) Chem Biol Drug Des 75: 51-67
- PubMed: 19954434 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1111/j.1747-0285.2009.00914.x
- Primary Citation Related Structures:
3KSL - PubMed Abstract:
Photoactive analogs of farnesyl diphosphate (FPP) are useful probes in studies of enzymes that employ this molecule as a substrate. Here, we describe the preparation and properties of two new FPP analogs that contain diazotrifluoropropanoyl photophores linked to geranyl diphosphate via amide or ester linkages. The amide-linked analog (3) was synthesized in 32P-labeled form from geraniol in seven steps. Experiments with Saccharomyces cerevisiae protein farnesyltransferase (ScPFTase) showed that 3 is an alternative substrate for the enzyme. Photolysis experiments with [(32)P]3 demonstrate that this compound labels the beta-subunits of both farnesyltransferase and geranylgeranyltransferase (types 1 and 2). However, the amide-linked probe 3 undergoes a rearrangement to a photochemically unreactive isomeric triazolone upon long term storage making it inconvenient to use. To address this stability issue, the ester-linked analog 4 was prepared in six steps from geraniol. Computational analysis and X-ray crystallographic studies suggest that 4 binds to protein farnesyl transferase (PFTase) in a similar fashion as FPP. Compound 4 is also an alternative substrate for PFTase, and a 32P-labeled form selectively photocrosslinks the beta-subunit of ScPFTase as well as E. coli farnesyldiphosphate synthase and a germacrene-producing sesquiterpene synthase from Nostoc sp. strain PCC7120 (a cyanobacterial source). Finally, nearly exclusive labeling of ScPFTase in crude E. coli extract was observed, suggesting that [32P]4 manifests significant selectivity and should hence be useful for identifying novel FPP-utilizing enzymes in crude protein preparations.
- Department of Chemistry, University of Minnesota, Minneapolis, MN 55455, USA.
Organizational Affiliation: