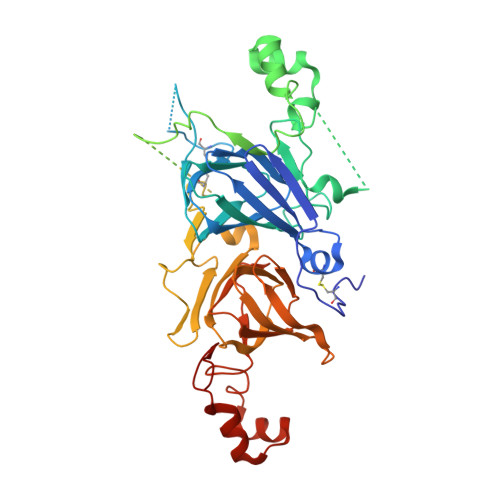
Conservation and divergence on plant seed 11S globulins based on crystal structures.
Tandang-Silvas, M.R., Fukuda, T., Fukuda, C., Prak, K., Cabanos, C., Kimura, A., Itoh, T., Mikami, B., Utsumi, S., Maruyama, N.(2010) Biochim Biophys Acta 1804: 1432-1442
- PubMed: 20215054 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbapap.2010.02.016
- Primary Citation Related Structures:
2D5F, 2D5H, 2E9Q, 3KGL, 3KSC - PubMed Abstract:
The crystal structures of two pro-11S globulins namely: rapeseed procruciferin and pea prolegumin are presented here. We have extensively compared them with the other known structures of plant seed 11S and 7S globulins. In general, the disordered regions in the crystal structures among the 11S globulins correspond to their five variable regions. Variable region III of procruciferin is relatively short and is in a loop conformation. This region is highly disordered in other pro-11S globulin crystals. Local helical and strand variations also occur across the group despite general structure conservation. We showed how these variations may alter specific physicochemical, functional and physiological properties. Aliphatic hydrophobic residues on the molecular surface correlate well with Tm values of the globulins. We also considered other structural features that were reported to influence thermal stability but no definite conclusion was drawn since each factor has additive or subtractive effect. Comparison between proA3B4 and mature A3B4 revealed an increase in r.m.s.d. values near variable regions II and IV. Both regions are on the IE face. Secondary structure based alignment of 11S and 7S globulins revealed 16 identical residues. Based on proA3B4 sequence, Pro60, Gly128, Phe163, Phe208, Leu213, Leu227, Ile237, Pro382, Val404, Pro425 and Val 466 are involved in trimer formation and stabilization. Gly28, Gly74, Asp135, Gly349 and Gly397 are involved in correct globular folding.
- Laboratory of Food Quality Design and Development, Graduate School of Agriculture, Kyoto University, Uji, Kyoto, Kyoto 611-0011, Japan.
Organizational Affiliation: