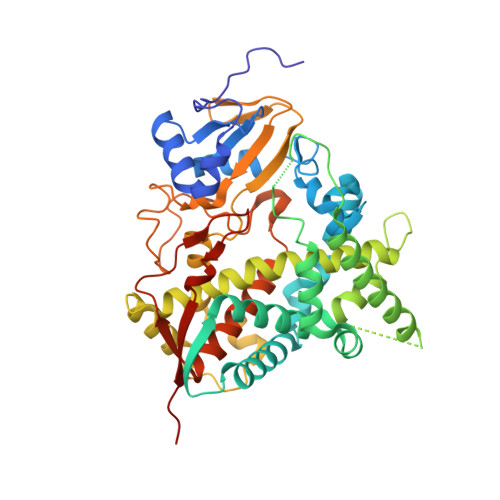
Structural Insights into Inhibition of Sterol 14{alpha}-Demethylase in the Human Pathogen Trypanosoma cruzi.
Lepesheva, G.I., Hargrove, T.Y., Anderson, S., Kleshchenko, Y., Furtak, V., Wawrzak, Z., Villalta, F., Waterman, M.R.(2010) J Biological Chem 285: 25582-25590
- PubMed: 20530488 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M110.133215
- Primary Citation Related Structures:
3K1O, 3KHM, 3KSW - PubMed Abstract:
Trypanosoma cruzi causes Chagas disease (American trypanosomiasis), which threatens the lives of millions of people and remains incurable in its chronic stage. The antifungal drug posaconazole that blocks sterol biosynthesis in the parasite is the only compound entering clinical trials for the chronic form of this infection. Crystal structures of the drug target enzyme, Trypanosoma cruzi sterol 14alpha-demethylase (CYP51), complexed with posaconazole, another antifungal agent fluconazole and an experimental inhibitor, (R)-4'-chloro-N-(1-(2,4-dichlorophenyl)-2-(1H-imid-azol-1-yl)ethyl)biphenyl-4-carboxamide (VNF), allow prediction of important chemical features that enhance the drug potencies. Combined with comparative analysis of inhibitor binding parameters, influence on the catalytic activity of the trypanosomal enzyme and its human counterpart, and their cellular effects at different stages of the Trypanosoma cruzi life cycle, the structural data provide a molecular background to CYP51 inhibition and azole resistance and enlighten the path for directed design of new, more potent and selective drugs to develop an efficient treatment for Chagas disease.
- Department of Biochemistry, School of Medicine, Vanderbilt University, Nashville, Tennessee 37232, USA. i.lepesheva@vanderbilt.edu
Organizational Affiliation: