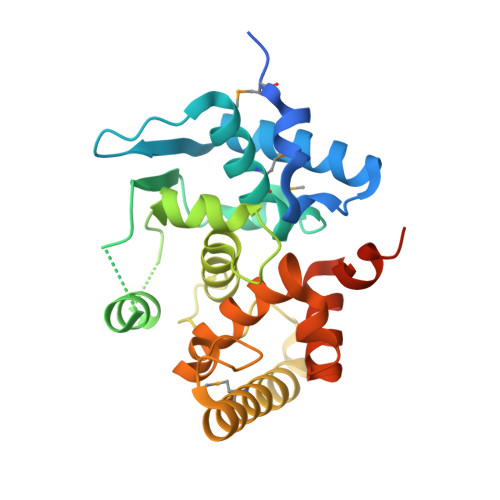
The Structure of Mlc Titration Factor A (MtfA/YeeI) Reveals a Prototypical Zinc Metallopeptidase Related to Anthrax Lethal Factor.
Xu, Q., Gohler, A.K., Kosfeld, A., Carlton, D., Chiu, H.J., Klock, H.E., Knuth, M.W., Miller, M.D., Elsliger, M.A., Deacon, A.M., Godzik, A., Lesley, S.A., Jahreis, K., Wilson, I.A.(2012) J Bacteriol 194: 2987-2999
- PubMed: 22467785 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/JB.00038-12
- Primary Citation Related Structures:
3DL1, 3KHI - PubMed Abstract:
MtfA of Escherichia coli (formerly YeeI) was previously identified as a regulator of the phosphoenolpyruvate (PEP)-dependent:glucose phosphotransferase system. MtfA homolog proteins are highly conserved, especially among beta- and gammaproteobacteria. We determined the crystal structures of the full-length MtfA apoenzyme from Klebsiella pneumoniae and its complex with zinc (holoenzyme) at 2.2 and 1.95 Å, respectively. MtfA contains a conserved H(149)E(150)XXH(153)+E(212)+Y(205) metallopeptidase motif. The presence of zinc in the active site induces significant conformational changes in the region around Tyr205 compared to the conformation of the apoenzyme. Additionally, the zinc-bound MtfA structure is in a self-inhibitory conformation where a region that was disordered in the unliganded structure is now observed in the active site and a nonproductive state of the enzyme is formed. MtfA is related to the catalytic domain of the anthrax lethal factor and the Mop protein involved in the virulence of Vibrio cholerae, with conservation in both overall structure and in the residues around the active site. These results clearly provide support for MtfA as a prototypical zinc metallopeptidase (gluzincin clan).
- Joint Center for Structural Genomics, SLAC National Accelerator Laboratory, Menlo Park, CA, USA.
Organizational Affiliation: