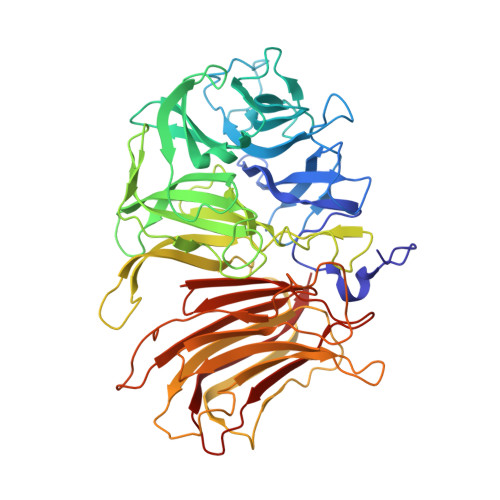
Structural and kinetic analysis of Schwanniomyces occidentalis invertase reveals a new oligomerization pattern and the role of its supplementary domain in substrate binding
Alvaro-Benito, M., Polo, A., Gonzalez, B., Fernandez-Lobato, M., Sanz-Aparicio, J.(2010) J Biological Chem 285: 13930-13941
- PubMed: 20181943 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M109.095430
- Primary Citation Related Structures:
3KF3, 3KF5 - PubMed Abstract:
Schwanniomyces occidentalis invertase is an extracellular enzyme that hydrolyzes sucrose and releases beta-fructose from various oligosaccharides and essential storage fructan polymers such as inulin. We report here the three-dimensional structure of Sw. occidentalis invertase at 2.9 A resolution and its complex with fructose at 1.9 A resolution. The monomer presents a bimodular arrangement common to other GH32 enzymes, with an N-terminal 5-fold beta-propeller catalytic domain and a C-terminal beta-sandwich domain for which the function has been unknown until now. However, the dimeric nature of Sw. occidentalis invertase reveals a unique active site cleft shaped by both subunits that may be representative of other yeast enzymes reported to be multimeric. Binding of the tetrasaccharide nystose and the polymer inulin was explored by docking analysis, which suggested that medium size and long substrates are recognized by residues from both subunits. The identified residues were mutated, and the enzymatic activity of the mutants against sucrose, nystose, and inulin were investigated by kinetic analysis. The replacements that showed the largest effect on catalytic efficiency were Q228V, a residue putatively involved in nystose and inulin binding, and S281I, involved in a polar link at the dimer interface. Moreover, a significant decrease in catalytic efficiency against inulin was observed in the mutants Q435A and Y462A, both located in the beta-sandwich domain of the second monomer. This highlights the essential function that oligomerization plays in substrate specificity and assigns, for the first time, a direct catalytic role to the supplementary domain of a GH32 enzyme.
- Centro de Biología Molecular Severo Ochoa, Departamento de Biología Molecular, Consejo Superior de Investigaciones Cientificas-Universidad Autónoma de Madrid, Cantoblanco, 28049 Madrid, Spain.
Organizational Affiliation: