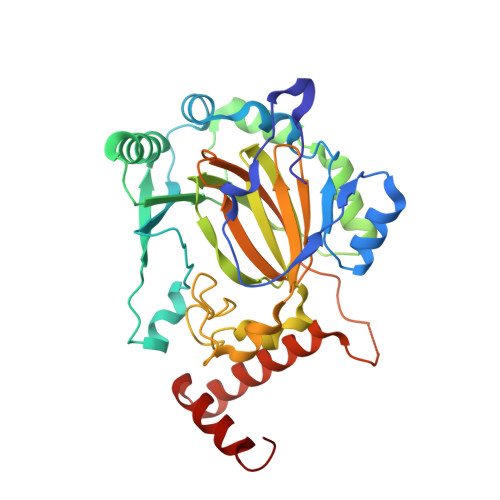
Crystal structures of human FIH-1 in complex with quinol family inhibitors
Moon, H., Han, S., Park, H., Choe, J.(2010) Mol Cells 29: 471-474
- PubMed: 20396966 Search on PubMed
- DOI: https://doi.org/10.1007/s10059-010-0058-3
- Primary Citation Related Structures:
3KCX, 3KCY - PubMed Abstract:
Hypoxia-Inducible Factor-1 (HIF-1) plays an important role as a transcription factor under hypoxia. It activates numerous genes including those involved in angiogenesis, glucose metabolisms, cell proliferation and cell survival. The HIF-1 alpha subunit is regulated by 2-oxoglutarate (OG)- and Fe(II)-dependent hydroxylases, including Factor Inhibiting HIF-1 (FIH-1). FIH-1 hydroxylates Asn803 of HIF-1 alpha and blocks its interaction with co-activating molecules. Quinol family compounds such as 5-chloro-7-iodo-8-hydroxyquinoline (Clioquinol) have been shown to inhibit the hydroxylation activity of FIH-1. Here we determined the complex crystal structures of FIH-1: Clioquinol and FIH-1: 8-Hydroxyquinoline. Clioquinol and 8-Hydroxyquinoline bind to the active site of FIH-1 by coordinating the Fe(II) ion, thereby inhibiting the binding of a co-substrate, 2OG. Contrary to other known FIH-1 inhibitors that have negative charges, Clioquinol and 8-hydroxyquinoline are neutral in charge and can provide a template for improved inhibitor design that can selectively inhibit FIH-1.
- Department of Life Science, University of Seoul, Seoul 130-743, Korea.
Organizational Affiliation: