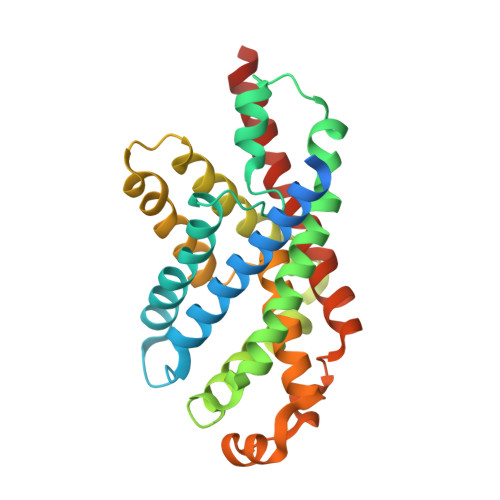
Structure of the formate transporter FocA reveals a pentameric aquaporin-like channel
Wang, Y., Huang, Y., Wang, J., Cheng, C., Huang, W., Lu, P., Xu, Y.-N., Wang, P., Yan, N., Shi, Y.(2009) Nature 462: 467-472
- PubMed: 19940917 Search on PubMed
- DOI: https://doi.org/10.1038/nature08610
- Primary Citation Related Structures:
3KCU, 3KCV - PubMed Abstract:
FocA is a representative member of the formate-nitrite transporter family, which transports short-chain acids in bacteria, archaea, fungi, algae and parasites. The structure and transport mechanism of the formate-nitrite transporter family remain unknown. Here we report the crystal structure of Escherichia coli FocA at 2.25 A resolution. FocA forms a symmetric pentamer, with each protomer consisting of six transmembrane segments. Despite a lack of sequence homology, the overall structure of the FocA protomer closely resembles that of aquaporin and strongly argues that FocA is a channel, rather than a transporter. Structural analysis identifies potentially important channel residues, defines the channel path and reveals two constriction sites. Unlike aquaporin, FocA is impermeable to water but allows the passage of formate. A structural and biochemical investigation provides mechanistic insights into the channel activity of FocA.
- Ministry of Education Protein Science Laboratory, Center for Structural Biology, School of Life Sciences and School of Medicine, Tsinghua University, Beijing 100084, China.
Organizational Affiliation: