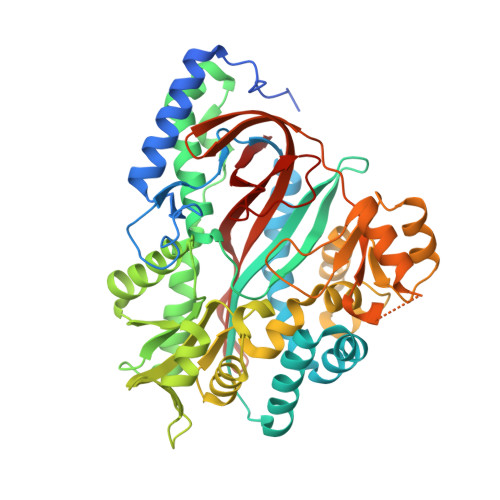
Structural Basis for Evolution of Product Diversity in Soybean Glutathione Biosynthesis.
Galant, A., Arkus, K.A., Zubieta, C., Cahoon, R.E., Jez, J.M.(2009) Plant Cell 21: 3450-3458
- PubMed: 19948790 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1105/tpc.109.071183
- Primary Citation Related Structures:
3KAJ, 3KAK, 3KAL - PubMed Abstract:
The redox active peptide glutathione is ubiquitous in nature, but some plants also synthesize glutathione analogs in response to environmental stresses. To understand the evolution of chemical diversity in the closely related enzymes homoglutathione synthetase (hGS) and glutathione synthetase (GS), we determined the structures of soybean (Glycine max) hGS in three states: apoenzyme, bound to gamma-glutamylcysteine (gammaEC), and with hGSH, ADP, and a sulfate ion bound in the active site. Domain movements and rearrangement of active site loops change the structure from an open active site form (apoenzyme and gammaEC complex) to a closed active site form (hGSH*ADP*SO(4)(2-) complex). The structure of hGS shows that two amino acid differences in an active site loop provide extra space to accommodate the longer beta-Ala moiety of hGSH in comparison to the glycinyl group of glutathione. Mutation of either Leu-487 or Pro-488 to an Ala improves catalytic efficiency using Gly, but a double mutation (L487A/P488A) is required to convert the substrate preference of hGS from beta-Ala to Gly. These structures, combined with site-directed mutagenesis, reveal the molecular changes that define the substrate preference of hGS, explain the product diversity within evolutionarily related GS-like enzymes, and reinforce the critical role of active site loops in the adaptation and diversification of enzyme function.
- Department of Biology, Washington University, St. Louis, Missouri 63130, USA.
Organizational Affiliation: